-Title: Next-generation file formats (NGFF)
-Shortname: ome-ngff
+Title: OME-Zarr specification
+Shortname: ome-zarr
Level: 1
Status: LS-COMMIT
Status: w3c/ED
@@ -16,6 +16,8 @@ Markup Shorthands: markdown yes
Editor: Josh Moore, University of Dundee (UoD) https://www.dundee.ac.uk, https://orcid.org/0000-0003-4028-811X
Editor: Sébastien Besson, University of Dundee (UoD) https://www.dundee.ac.uk, https://orcid.org/0000-0001-8783-1429
Editor: Constantin Pape, European Molecular Biology Laboratory (EMBL) https://www.embl.org/sites/heidelberg/, https://orcid.org/0000-0001-6562-7187
+Editor: John Bogovic, Hughes Medical Institute Janelia (HHMI) https://www.janelia.org/, https://orcid.org/0000-0002-4829-9457
+Editor: Norman Rzepka, scalable minds https://scalableminds.com/, https://orcid.org/0000-0002-8168-7929
Text Macro: NGFFVERSION 0.5-dev
Abstract: This document contains next-generation file format (NGFF)
Abstract: specifications for storing bioimaging data in the cloud.
@@ -122,11 +124,13 @@ be stored on a web server to be accessed via HTTP or in object storage
like S3 or GCS.
OME-Zarr is an implementation of the OME-NGFF specification using the Zarr
-format. Arrays MUST be defined and stored in a hierarchical organization as
-defined by the
-[version 2 of the Zarr specification ](https://zarr.readthedocs.io/en/stable/spec/v2.html).
-OME-NGFF metadata MUST be stored as attributes in the corresponding Zarr
-groups.
+format. Arrays and groups MUST be defined and stored in a hierarchical organization as
+defined by the [version 3 of the Zarr specification](https://zarr.readthedocs.io/en/stable/spec/v3.html).
+OME metadata MUST be stored as attributes in the corresponding Zarr groups.
+
+All features of the Zarr format as finalized through the [ZEP process](https://zarr.dev/zeps/active/ZEP0000.html) are available in OME-Zarr.
+This explicitly includes chunk key encodings, codecs, data types, and storage transformers.
+Restrictions may be introduced in the future.
Images {#image-layout}
----------------------
@@ -144,39 +148,45 @@ For this example we assume an image with 5 dimensions and axes called `t,c,z,y,x
│
└── 456.zarr # Another image (id=456) converted to Zarr.
│
- ├── .zgroup # Each image is a Zarr group, or a folder, of other groups and arrays.
- ├── .zattrs # Group level attributes are stored in the .zattrs file and include
- │ # "multiscales" and "omero" (see below). In addition, the group level attributes
- │ # must also contain "_ARRAY_DIMENSIONS" if this group directly contains multi-scale arrays.
+ ├── zarr.json # Each image is a Zarr group, or a folder, of other groups and arrays.
+ │ # Group level attributes are stored in the zarr.json file and include
+ │ # "multiscales" and "omero" (see below).
│
├── 0 # Each multiscale level is stored as a separate Zarr array,
│ ... # which is a folder containing chunk files which compose the array.
├── n # The name of the array is arbitrary with the ordering defined by
│ │ # by the "multiscales" metadata, but is often a sequence starting at 0.
│ │
- │ ├── .zarray # All image arrays must be up to 5-dimensional
+ │ ├── zarr.json # All image arrays must be up to 5-dimensional
│ │ # with the axis of type time before type channel, before spatial axes.
│ │
- │ └─ t # Chunks are stored with the nested directory layout.
- │ └─ c # All but the last chunk element are stored as directories.
- │ └─ z # The terminal chunk is a file. Together the directory and file names
- │ └─ y # provide the "chunk coordinate" (t, c, z, y, x), where the maximum coordinate
- │ └─ x # will be `dimension_size / chunk_size`.
+ │ └─ c # Chunks are stored with the chunk key encoding as specified in the zarr.json.
+ | | # In this example, the chunk key encoding is "default" with separator "/".
+ | | # The chunk keys (composed of file path and file name) provide the "chunk coordinate" (t, c, z, y, x).
+ │ ├─ 0 # Time dimension
+ | | ├─ 0 # Channel dimension
+ | | | ├─ 0 # Z dimension
+ | | | | ├─ 0 # Y dimension
+ | | | | | ├─ 0 # X dimension
+ | | | | | └─ ...
+ | | | | └─ ...
+ | | | └─ ...
+ | | └─ ...
+ │ └─ ...
│
└── labels
│
- ├── .zgroup # The labels group is a container which holds a list of labels to make the objects easily discoverable
- │
- ├── .zattrs # All labels will be listed in `.zattrs` e.g. `{ "labels": [ "original/0" ] }`
+ ├── zarr.json # The labels group is a container which holds one or more multiscale labels.
+ │ # All labels will be listed in the top-level attributes e.g. `{ "labels": [ "labels/original/0" ] }`
│ # Each dimension of the label `(t, c, z, y, x)` should be either the same as the
│ # corresponding dimension of the image, or `1` if that dimension of the label
│ # is irrelevant.
│
└── original # Intermediate folders are permitted but not necessary and currently contain no extra metadata.
│
- └── 0 # Multiscale, labeled image. The name is unimportant but is registered in the "labels" group above.
- ├── .zgroup # Zarr Group which is both a multiscaled image as well as a labeled image.
- ├── .zattrs # Metadata of the related image and as well as display information under the "image-label" key.
+ └── 0 # Multiscale, labeled image. The name is unimportant but is registered in the "labels" metadata above.
+ ├── zarr.json # Zarr Group which is both a multiscaled image as well as a labeled image including
+ │ # metadata of the related image and as well as display information under the "image-label" key.
│
├── 0 # Each multiscale level is stored as a separate Zarr array, as above, but only integer values
│ ... # are supported.
@@ -189,7 +199,7 @@ High-content screening {#hcs-layout}
------------------------------------
The following specification defines the hierarchy for a high-content screening
-dataset. Three groups MUST be defined above the images:
+dataset. Three groups must be defined above the images:
- the group above the images defines the well and MUST implement the
[well specification](#well-md). All images contained in a well are fields
@@ -199,27 +209,21 @@ dataset. Three groups MUST be defined above the images:
collection of wells organized in rows and columns. It MUST implement the
[plate specification](#plate-md)
-A well row group SHOULD NOT be present if there are no images in the well row.
-A well group SHOULD NOT be present if there are no images in the well.
-
. # Root folder, potentially in S3,
│
└── 5966.zarr # One plate (id=5966) converted to Zarr
- ├── .zgroup
- ├── .zattrs # Implements "plate" specification
+ ├── zarr.json # Implements "plate" specification
├── A # First row of the plate
- │ ├── .zgroup
+ │ ├── zarr.json
│ │
│ ├── 1 # First column of row A
- │ │ ├── .zgroup
- │ │ ├── .zattrs # Implements "well" specification
+ │ │ ├── zarr.json # Implements "well" specification
│ │ │
│ │ ├── 0 # First field of view of well A1
│ │ │ │
- │ │ │ ├── .zgroup
- │ │ │ ├── .zattrs # Implements "multiscales", "omero"
+ │ │ │ ├── zarr.json # Implements "multiscales", "omero"
│ │ │ ├── 0
│ │ │ │ ... # Resolution levels
│ │ │ ├── n
@@ -235,20 +239,25 @@ A well group SHOULD NOT be present if there are no images in the well.
Metadata {#metadata}
====================
-The various `.zattrs` files throughout the above array hierarchy may contain metadata
+The various `zarr.json` files throughout the above array hierarchy may contain OME metadata
keys as specified below for discovering certain types of data, especially images.
+All OME metadata is stored under a `ome` key within the `attributes` of these `zarr.json` files.
-"axes" metadata {#axes-md}
---------------------------
+
-"axes" describes the dimensions of a physical coordinate space. It is a list of dictionaries, where each dictionary describes a dimension (axis) and:
-- MUST contain the field "name" that gives the name for this dimension. The values MUST be unique across all "name" fields.
-- SHOULD contain the field "type". It SHOULD be one of "space", "time" or "channel", but MAY take other values for custom axis types that are not part of this specification yet.
-- SHOULD contain the field "unit" to specify the physical unit of this dimension. The value SHOULD be one of the following strings, which are valid units according to UDUNITS-2.
- - Units for "space" axes: 'angstrom', 'attometer', 'centimeter', 'decimeter', 'exameter', 'femtometer', 'foot', 'gigameter', 'hectometer', 'inch', 'kilometer', 'megameter', 'meter', 'micrometer', 'mile', 'millimeter', 'nanometer', 'parsec', 'petameter', 'picometer', 'terameter', 'yard', 'yoctometer', 'yottameter', 'zeptometer', 'zettameter'
- - Units for "time" axes: 'attosecond', 'centisecond', 'day', 'decisecond', 'exasecond', 'femtosecond', 'gigasecond', 'hectosecond', 'hour', 'kilosecond', 'megasecond', 'microsecond', 'millisecond', 'minute', 'nanosecond', 'petasecond', 'picosecond', 'second', 'terasecond', 'yoctosecond', 'yottasecond', 'zeptosecond', 'zettasecond'
-
-If part of [[#multiscale-md]], the length of "axes" MUST be equal to the number of dimensions of the arrays that contain the image data.
+```json
+{
+ "zarr_format": 3,
+ "node_type": "group",
+ "attributes": {
+ "ome": {
+ "version": "0.5-dev",
+ ...
+ }
+ }
+}
+```
+
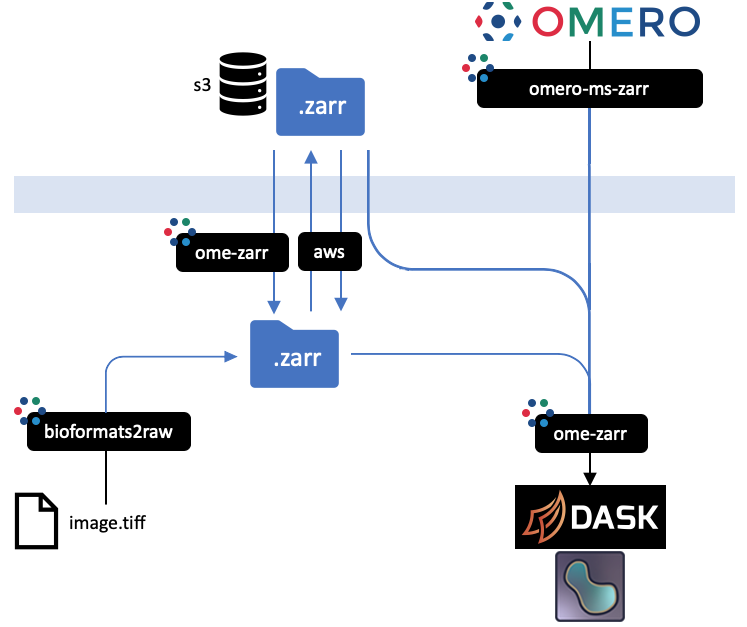
"bioformats2raw.layout" (transitional) {#bf2raw}
------------------------------------------------
@@ -267,11 +276,9 @@ Typical Zarr layout produced by running `bioformats2raw` on a fileset that conta
series.ome.zarr # One converted fileset from bioformats2raw
- ├── .zgroup
- ├── .zattrs # Contains "bioformats2raw.layout" metadata
+ ├── zarr.json # Contains "bioformats2raw.layout" metadata
├── OME # Special group for containing OME metadata
- │ ├── .zgroup
- │ ├── .zattrs # Contains "series" metadata
+ │ ├── zarr.json # Contains "series" metadata
│ └── METADATA.ome.xml # OME-XML file stored within the Zarr fileset
├── 0 # First image in the collection
├── 1 # Second image in the collection
@@ -280,7 +287,7 @@ series.ome.zarr # One converted fileset from bioformats2raw
Attributes
-The top-level `.zattrs` file must contain the `bioformats2raw.layout` key:
+The top-level OME group attributes must contain the `bioformats2raw.layout` key:
path: examples/bf2raw/image.json
highlight: json
@@ -295,7 +302,7 @@ path: examples/bf2raw/plate.json
highlight: json
-The `.zattrs` file within the OME group may contain the "series" key:
+The attributes within the OME group may contain the "series" key:
path: examples/ome/series-2.json
@@ -306,7 +313,7 @@ highlight: json
Conforming groups:
-- MUST have the value "3" for the "bioformats2raw.layout" key in their `.zattrs` metadata at the top of the hierarchy;
+- MUST have the value "3" for the "bioformats2raw.layout" key in their OME attribute metadata at the top of the hierarchy;
- SHOULD have OME metadata representing the entire collection of images in a file named "OME/METADATA.ome.xml" which:
- MUST adhere to the OME-XML specification but
- MUST use `
+path: examples/coordSystems/arrayCoordSys.json
+highlight: json
+
+
+
+
+
+### Coordinate convention
+
+**The pixel/voxel center is the origin of the continuous coordinate system.**
+
+It is vital to consistently define relationship between the discrete/array and continuous/interpolated
+coordinate systems. A pixel/voxel is the continuous region (rectangle) that corresponds to a single sample
+in the discrete array, i.e., the area corresponding to nearest-neighbor (NN) interpolation of that sample.
+The center of a 2d pixel corresponding to the origin `(0,0)` in the discrete array is the origin of the continuous coordinate
+system `(0.0, 0.0)` (when the transformation is the identity). The continuous rectangle of the pixel is given by the
+half-open interval `[-0.5, 0.5) x [-0.5, 0.5)` (i.e., -0.5 is included, +0.5 is excluded). See chapter 4 and figure 4.1 of the ITK Software Guide [[itk]].
+=======
+
+
+
"coordinateTransformations" metadata {#trafo-md}
------------------------------------------------
-"coordinateTransformations" describe a series of transformations that map between two coordinate spaces (defined by "axes").
-For example, to map a discrete data space of an array to the corresponding physical space.
-It is a list of dictionaries. Each entry describes a single transformation and MUST contain the field "type".
-The value of "type" MUST be one of the elements of the `type` column in the table below.
-Additional fields for the entry depend on "type" and are defined by the column `fields`.
+"coordinateTransformations" describe the mapping between two coordinate systems (defined by "axes").
+For example, to map an array's discrete coordinate system to its corresponding physical coordinates.
+Coordinate transforms are in the "forward" direction. They represent functions from *points* in the
+input space to *points* in the output space.
+
+
+- MUST contain the field "type".
+- MUST contain any other fields required by the given "type" (see table below).
+- MUST contain the field "output", unless part of a `sequence` or `inverseOf` (see details).
+- MUST contain the field "input", unless part of a `sequence` or `inverseOf` (see details).
+- MAY contain the field "name". Its value MUST be unique across all "name" fields for coordinate transformations.
+- Parameter values MUST be compatible with input and output space dimensionality (see details).
| `identity` | identity transformation, is the default transformation and is typically not explicitly defined - | ||||
|---|---|---|---|---|---|
| `translation` | one of: `"translation":List[float]`, `"path":str` | translation vector, stored either as a list of floats (`"translation"`) or as binary data at a location in this container (`path`). The length of vector defines number of dimensions. | - | |||
| `scale` | one of: `"scale":List[float]`, `"path":str` | scale vector, stored either as a list of floats (`scale`) or as binary data at a location in this container (`path`). The length of vector defines number of dimensions. | + | |||
| `identity` + | + | The identity transformation is the default transformation and is typically not explicitly defined. + | |||
| `mapIndex` + | `"mapIndex":List[number]` + | A `mapIndex` transformation specifies an axis permutation by reordering the input axes. + | |||
| `mapAxis` + | `"mapAxis":Dict[String:String]` + | A `maxAxis` transformation specifies an axis permutation as a map between axis names. + | |||
| `translation` + | one of: `"translation":List[number]`, `"path":str` + | translation vector, stored either as a list of numbers (`"translation"`) or as binary data at a location + in this container (`path`). + | |||
| `scale` + | one of: `"scale":List[number]`, `"path":str` + | scale vector, stored either as a list of numbers (`scale`) or as binary data at a location in this + container (`path`). + | |||
| `affine` + | one of: `"affine":List[number]`, `"path":str` + | affine transformation matrix stored as a flat array stored either with json uing the affine field + or as binary data at a location in this container (path). If both are present, the binary values at path should be used. + | |||
| `rotation` + | one of: `"rotation":List[number]`, `"path":str` + | rotation transformation matrix stored as an array stored either + with json or as binary data at a location in this container (path). + If both are present, the binary parameters at path are used. + | |||
| `sequence` + | `"transformations":List[Transformation]` + | A sequence of transformations, Applying the sequence applies the composition of all transforms in the list, in order. + | |||
| `displacements` + | `"path":str` `"interpolation":str` + | Displacement field transformation located at (path). + | |||
| `coordinates` + | `"path":str` `"interpolation":str` + | Coordinate field transformation located at (path). + | |||
| `inverseOf` + | `"transform":Transform` + | The inverse of a transformation. Useful if a transform is not closed-form invertible. See Forward and inverse for details and examples. + | |||
| `bijection` + | `"forward":Transform` `"inverse":Transform` + | Explicitly define an invertible transformation by providing a forward transformation and its inverse. + | |||
| `byDimension` + | `"transformations":List[Transformation]` + | Define a high dimensional transformation using lower dimensional transformations on subsets of
+ dimensions.
| type | fields | description
| |
+store.zarr # Root folder of the zarr store +│ +├── zarr.json # coordinate transformations describing the relationship between two image coordinate systems +│ # are stored in the attributes of their parent group. +│ # transformations between 'volume' and 'crop' coordinate systems are stored here. +│ +├── coordinateTransformations # transformations that use array storage go in a "transformations" zarr group. +│ └── displacements # for example, a zarray containing a displacement field +│ └── zarr.json +│ +├── volume +│ ├── zarr.json # group level attributes (multiscales) +│ └── 0 # a group containing the 0th scale +│ └── image # a zarr array +│ └── zarr.json # physical coordinate system and transformations here the array attributes +└── crop + ├── zarr.json # group level attributes (multiscales) + └── 0 # a group containing the 0th scale + └── image # a zarr array + └── zarr.json # physical coordinate system and transformations here the array attributes ++ +### Additional details + +Most coordinate transformations MUST specify their input and output coordinate systems using `input` and `output` with a string value +corresponding to the name of a coordinate system. The coordinate system's name may be the path to an array, and therefore may +not appear in the list of coordinate systems. + +Exceptions are if the the coordinate transformation appears in the `transformations` list of a `sequence` or is the +`transformation` of an `inverseOf` transformation. In these two cases input and output SHOULD be omitted (see below for +details). + +Transformations in the `transformations` list of a `byDimensions` transforemation MUST provide `input` and `output` as arrays +of strings corresponding to axis names of the parent transformation's input and output coordinate systems (see below for +details). + +
+path: examples/transformations/identity.json +highlight: json ++ +defines the function: + +``` +x = i +y = j +``` + +
+path: examples/transformations/mapIndex1.json +highlight: json ++ +The "equivalent to identity" transformation defines the function: + +``` +x = i +y = j +``` + +and the "permutation" transformation defines the function + +``` +x = j +y = i +``` + +
+path: examples/transformations/mapIndex2.json +highlight: json ++ +The "projection_down" transformation defines the function: + +``` +x = b +``` + +and the "projection_up" transformation defines the function: + +``` +x = a +y = b +z = b +``` + +
+path: examples/transformations/mapAxis1.json +highlight: json ++ +The "equivalent to identity" transformation defines the function: + +``` +x = i +y = j +``` + +and the "permutation" transformation defines the function + +``` +x = j +y = i +``` + +
+path: examples/transformations/mapAxis2.json +highlight: json ++ +The "projection_down" transformation defines the function: + +``` +x = b +``` + +and the "projection_up" transformation defines the function: + +``` +x = a +y = b +z = b +``` +
+path: examples/transformations/translation.json +highlight: json ++ +defines the function: + +``` +x = i + 9 +y = j - 1.42 +``` +
+path: examples/transformations/scale.json +highlight: json ++ +defines the function: + +``` +x = 3.12 * i +y = 2 * j +``` +
+ path: examples/transformations/affine2d2d.json + highlight: json ++ + defines the function: + + ``` + x = 1*i + 2*j + 3 + y = 4*i + 5*j + 6 + ``` +
+ path: examples/transformations/affine2d3d.json + highlight: json ++ + defines the function: + + ``` + x = 1*i + 2*j + 3 + y = 4*i + 5*j + 6 + z = 7*i + 8*j + 9 + ``` + + Using a nested 2D JSON array, the same transformation can be written: +
+ path: examples/transformations/affine2d3d_nested.json + highlight: json ++ +
+ path: examples/transformations/rotation.json + highlight: json ++ + defines the function: + + ``` + x = 0*i - 1*j + y = 1*i + 0*j + ``` +
+ path: examples/transformations/inverseOf.json + highlight: json ++ +
+path: examples/transformations/sequence.json +highlight: json ++ +describes the function + +``` +x = (i + 0.1) * 2 +y = (j + 0.9) * 3 +``` + +and is invertible. +
linear (default)nearestcubic+path: examples/transformations/byDimension1.json +highlight: json ++ +
+path: examples/transformations/byDimension2.json +highlight: json ++ +
+path: examples/transformations/byDimensionInvalid1.json +highlight: json ++ +It is invalid for two reasons. First because input `0` used by the scale transformation is not an axis of the `byDimension` transformation's `input`. Second, the `x` axis of the `output` does not appear in the `output` of any child transformation. + +
+path: examples/transformations/byDimensionInvalid2.json +highlight: json ++ +This transformation is invalid because the output axis `x` appears in more than one transformation in the `transformations` list. + +
+path: examples/transformations/bijection.json +highlight: json ++ +the input and output of the `forward` and `inverse` transformations are understoood to be: + +
+path: examples/transformations/bijection_verbose.json +highlight: json ++ +
 diff --git a/latest/schemas/bf2raw.schema b/latest/schemas/bf2raw.schema
index 834aee24..d661c8bf 100644
--- a/latest/schemas/bf2raw.schema
+++ b/latest/schemas/bf2raw.schema
@@ -1,14 +1,16 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/bf2raw.schema",
- "title": "NGFF container produced by bioformats2raw",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "Zarr container produced by bioformats2raw",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
"bioformats2raw.layout": {
"description": "The top-level identifier metadata added by bioformats2raw",
"type": "number",
- "enum": [3]
+ "enum": [
+ 3
+ ]
}
}
}
diff --git a/latest/schemas/image.schema b/latest/schemas/image.schema
index 3926c4f4..79f64f99 100644
--- a/latest/schemas/image.schema
+++ b/latest/schemas/image.schema
@@ -1,113 +1,136 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/image.schema",
- "title": "NGFF Image",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr Image",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
- "multiscales": {
- "description": "The multiscale datasets for this image",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "name": {
- "type": "string"
- },
- "datasets": {
- "type": "array",
- "minItems": 1,
- "items": {
- "type": "object",
- "properties": {
- "path": {
- "type": "string"
- },
- "coordinateTransformations": {
- "$ref": "#/$defs/coordinateTransformations"
- }
- },
- "required": ["path", "coordinateTransformations"]
- }
- },
- "version": {
- "type": "string",
- "enum": [
- "0.5-dev"
- ]
- },
- "axes": {
- "$ref": "#/$defs/axes"
- },
- "coordinateTransformations": {
- "$ref": "#/$defs/coordinateTransformations"
- }
- },
- "required": [
- "datasets", "axes"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "omero": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
"properties": {
- "channels": {
+ "version": {
+ "type": "string",
+ "enum": [
+ "0.5-dev"
+ ]
+ },
+ "multiscales": {
+ "description": "The multiscale datasets for this image",
"type": "array",
"items": {
"type": "object",
"properties": {
- "window": {
+ "name": {
+ "type": "string"
+ },
+ "datasets": {
+ "type": "array",
+ "minItems": 1,
+ "items": {
+ "type": "object",
+ "properties": {
+ "path": {
+ "type": "string"
+ },
+ "coordinateTransformations": {
+ "$ref": "#/$defs/coordinateTransformations"
+ }
+ },
+ "required": [
+ "path",
+ "coordinateTransformations"
+ ]
+ }
+ },
+ "axes": {
+ "$ref": "#/$defs/axes"
+ },
+ "coordinateTransformations": {
+ "$ref": "#/$defs/coordinateTransformations"
+ },
+ "labels": {
+ "type": "array",
+ "items": {
+ "type": "string"
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ }
+ },
+ "required": [
+ "datasets",
+ "axes"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "omero": {
+ "type": "object",
+ "properties": {
+ "channels": {
+ "type": "array",
+ "items": {
"type": "object",
"properties": {
- "end": {
- "type": "number"
+ "window": {
+ "type": "object",
+ "properties": {
+ "end": {
+ "type": "number"
+ },
+ "max": {
+ "type": "number"
+ },
+ "min": {
+ "type": "number"
+ },
+ "start": {
+ "type": "number"
+ }
+ },
+ "required": [
+ "start",
+ "min",
+ "end",
+ "max"
+ ]
+ },
+ "label": {
+ "type": "string"
},
- "max": {
- "type": "number"
+ "family": {
+ "type": "string"
},
- "min": {
- "type": "number"
+ "color": {
+ "type": "string"
},
- "start": {
- "type": "number"
+ "active": {
+ "type": "boolean"
}
},
"required": [
- "start",
- "min",
- "end",
- "max"
+ "window",
+ "color"
]
- },
- "label": {
- "type": "string"
- },
- "family": {
- "type": "string"
- },
- "color": {
- "type": "string"
- },
- "active": {
- "type": "boolean"
}
- },
- "required": [
- "window",
- "color"
- ]
- }
+ }
+ },
+ "required": [
+ "channels"
+ ]
}
},
"required": [
- "channels"
+ "multiscales",
+ "version"
]
- }
+ },
+ "required": [
+ "ome"
+ ]
},
- "required": [ "multiscales" ],
-
"$defs": {
"axes": {
"type": "array",
@@ -122,7 +145,9 @@
},
"type": {
"type": "string",
- "enum": ["space"]
+ "enum": [
+ "space"
+ ]
},
"units": {
"type": "string"
@@ -141,10 +166,17 @@
},
"type": {
"type": "string",
- "enum": ["channel", "time", "space"]
+ "enum": [
+ "channel",
+ "time",
+ "space"
+ ]
}
},
- "required": ["name", "type"]
+ "required": [
+ "name",
+ "type"
+ ]
},
{
"type": "object",
@@ -155,11 +187,17 @@
"type": {
"type": "string",
"not": {
- "enum": ["space", "time", "channel"]
+ "enum": [
+ "space",
+ "time",
+ "channel"
+ ]
}
}
},
- "required": ["name"]
+ "required": [
+ "name"
+ ]
}
]
}
@@ -205,7 +243,10 @@
}
}
},
- "required": ["type", "scale"]
+ "required": [
+ "type",
+ "scale"
+ ]
},
{
"type": "object",
@@ -224,7 +265,10 @@
}
}
},
- "required": ["type", "translation"]
+ "required": [
+ "type",
+ "translation"
+ ]
}
]
}
diff --git a/latest/schemas/label.schema b/latest/schemas/label.schema
index 8b23d577..ebca25c4 100644
--- a/latest/schemas/label.schema
+++ b/latest/schemas/label.schema
@@ -1,75 +1,87 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/label.schema",
- "title": "OME-NGFF labelled image schema",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr labelled image schema",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
+ "required": [
+ "ome"
+ ],
"properties": {
- "image-label": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
+ "required": [
+ "version"
+ ],
"properties": {
- "colors": {
- "description": "The colors for this label image",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "label-value": {
- "description": "The value of the label",
- "type": "number"
- },
- "rgba": {
- "description": "The RGBA color stored as an array of four integers between 0 and 255",
- "type": "array",
- "items": {
- "type": "integer",
- "minimum": 0,
- "maximum": 255
- },
- "minItems": 4,
- "maxItems": 4
- }
- },
- "required": [
- "label-value"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "properties": {
- "description": "The properties for this label image",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "label-value": {
- "description": "The pixel value for this label",
- "type": "integer"
- }
- },
- "required": [
- "label-value"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "source": {
- "description": "The source of this label image",
- "type": "object",
- "properties": {
- "image": {
- "type": "string"
- }
- }
- },
"version": {
"description": "The version of the specification",
"type": "string",
"enum": [
"0.5-dev"
]
+ },
+ "image-label": {
+ "type": "object",
+ "properties": {
+ "colors": {
+ "description": "The colors for this label image",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "label-value": {
+ "description": "The value of the label",
+ "type": "number"
+ },
+ "rgba": {
+ "description": "The RGBA color stored as an array of four integers between 0 and 255",
+ "type": "array",
+ "items": {
+ "type": "integer",
+ "minimum": 0,
+ "maximum": 255
+ },
+ "minItems": 4,
+ "maxItems": 4
+ }
+ },
+ "required": [
+ "label-value"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "properties": {
+ "description": "The properties for this label image",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "label-value": {
+ "description": "The pixel value for this label",
+ "type": "integer"
+ }
+ },
+ "required": [
+ "label-value"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "source": {
+ "description": "The source of this label image",
+ "type": "object",
+ "properties": {
+ "image": {
+ "type": "string"
+ }
+ }
+ }
+ }
}
}
}
diff --git a/latest/schemas/ome.schema b/latest/schemas/ome.schema
index bd600a2a..dd4872e8 100644
--- a/latest/schemas/ome.schema
+++ b/latest/schemas/ome.schema
@@ -1,8 +1,8 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/ome.schema",
- "title": "NGFF group produced by bioformats2raw to contain OME metadata",
- "description": "JSON from OME-NGFF OME/.zattrs linked to an OME-XML file",
+ "title": "Zarr group produced by bioformats2raw to contain OME metadata",
+ "description": "JSON from OME-Zarr OME/zarr.json attributes linked to an OME-XML file",
"type": "object",
"properties": {
"series": {
diff --git a/latest/schemas/plate.schema b/latest/schemas/plate.schema
index 099f5bb4..150605e8 100644
--- a/latest/schemas/plate.schema
+++ b/latest/schemas/plate.schema
@@ -1,140 +1,156 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/0.4/schemas/plate.schema",
- "title": "OME-NGFF plate schema",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr plate schema",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
- "plate": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
"properties": {
- "acquisitions": {
- "description": "The acquisitions for this plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "id": {
- "description": "A unique identifier within the context of the plate",
- "type": "integer",
- "minimum": 0
- },
- "maximumfieldcount": {
- "description": "The maximum number of fields of view for the acquisition",
- "type": "integer",
- "exclusiveMinimum": 0
- },
- "name": {
- "description": "The name of the acquisition",
- "type": "string"
- },
- "description": {
- "description": "The description of the acquisition",
- "type": "string"
- },
- "starttime": {
- "description": "The start timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
- "type": "integer",
- "minimum": 0
- },
- "endtime": {
- "description": "The end timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
- "type": "integer",
- "minimum": 0
- }
- },
- "required": [
- "id"
- ]
- }
- },
"version": {
- "description": "The version of the specification",
"type": "string",
"enum": [
"0.5-dev"
]
},
- "field_count": {
- "description": "The maximum number of fields per view across all wells",
- "type": "integer",
- "exclusiveMinimum": 0
- },
- "name": {
- "description": "The name of the plate",
- "type": "string"
- },
- "columns": {
- "description": "The columns of the plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "name": {
- "description": "The column name",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+$"
+ "plate": {
+ "type": "object",
+ "properties": {
+ "acquisitions": {
+ "description": "The acquisitions for this plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "id": {
+ "description": "A unique identifier within the context of the plate",
+ "type": "integer",
+ "minimum": 0
+ },
+ "maximumfieldcount": {
+ "description": "The maximum number of fields of view for the acquisition",
+ "type": "integer",
+ "exclusiveMinimum": 0
+ },
+ "name": {
+ "description": "The name of the acquisition",
+ "type": "string"
+ },
+ "description": {
+ "description": "The description of the acquisition",
+ "type": "string"
+ },
+ "starttime": {
+ "description": "The start timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
+ "type": "integer",
+ "minimum": 0
+ },
+ "endtime": {
+ "description": "The end timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
+ "type": "integer",
+ "minimum": 0
+ }
+ },
+ "required": [
+ "id"
+ ]
}
},
- "required": [
- "name"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "rows": {
- "description": "The rows of the plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "name": {
- "description": "The row name",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+$"
- }
+ "field_count": {
+ "description": "The maximum number of fields per view across all wells",
+ "type": "integer",
+ "exclusiveMinimum": 0
},
- "required": [
- "name"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "wells": {
- "description": "The wells of the plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "path": {
- "description": "The path to the well subgroup",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+/[A-Za-z0-9]+$"
+ "name": {
+ "description": "The name of the plate",
+ "type": "string"
+ },
+ "columns": {
+ "description": "The columns of the plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "name": {
+ "description": "The column name",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+$"
+ }
+ },
+ "required": [
+ "name"
+ ]
},
- "rowIndex": {
- "description": "The index of the well in the rows list",
- "type": "integer",
- "minimum": 0
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "rows": {
+ "description": "The rows of the plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "name": {
+ "description": "The row name",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+$"
+ }
+ },
+ "required": [
+ "name"
+ ]
},
- "columnIndex": {
- "description": "The index of the well in the columns list",
- "type": "integer",
- "minimum": 0
- }
+ "minItems": 1,
+ "uniqueItems": true
},
- "required": [
- "path", "rowIndex", "columnIndex"
- ]
+ "wells": {
+ "description": "The wells of the plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "path": {
+ "description": "The path to the well subgroup",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+/[A-Za-z0-9]+$"
+ },
+ "rowIndex": {
+ "description": "The index of the well in the rows list",
+ "type": "integer",
+ "minimum": 0
+ },
+ "columnIndex": {
+ "description": "The index of the well in the columns list",
+ "type": "integer",
+ "minimum": 0
+ }
+ },
+ "required": [
+ "path",
+ "rowIndex",
+ "columnIndex"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ }
},
- "minItems": 1,
- "uniqueItems": true
+ "required": [
+ "columns",
+ "rows",
+ "wells"
+ ]
}
},
"required": [
- "columns", "rows", "wells"
+ "version",
+ "plate"
]
}
- }
+ },
+ "required": [
+ "ome"
+ ]
}
diff --git a/latest/schemas/strict_image.schema b/latest/schemas/strict_image.schema
index bcecc003..3499be75 100644
--- a/latest/schemas/strict_image.schema
+++ b/latest/schemas/strict_image.schema
@@ -9,11 +9,13 @@
"multiscales": {
"items": {
"required": [
- "version", "metadata", "type", "name"
+ "metadata",
+ "type",
+ "name"
]
}
}
}
}
]
-}
\ No newline at end of file
+}
diff --git a/latest/schemas/strict_label.schema b/latest/schemas/strict_label.schema
index c52dd51b..9d888276 100644
--- a/latest/schemas/strict_label.schema
+++ b/latest/schemas/strict_label.schema
@@ -8,7 +8,6 @@
"properties": {
"image-label": {
"required": [
- "version",
"colors"
]
}
diff --git a/latest/schemas/strict_plate.schema b/latest/schemas/strict_plate.schema
index 5a88ab72..4c68707f 100644
--- a/latest/schemas/strict_plate.schema
+++ b/latest/schemas/strict_plate.schema
@@ -18,8 +18,7 @@
}
},
"required": [
- "name",
- "version"
+ "name"
]
}
}
diff --git a/latest/schemas/strict_well.schema b/latest/schemas/strict_well.schema
index 1e200294..e70dff9f 100644
--- a/latest/schemas/strict_well.schema
+++ b/latest/schemas/strict_well.schema
@@ -7,9 +7,7 @@
{
"properties": {
"well": {
- "required": [
- "version"
- ]
+ "required": []
}
}
}
diff --git a/latest/schemas/well.schema b/latest/schemas/well.schema
index 891abd05..651b8dc0 100644
--- a/latest/schemas/well.schema
+++ b/latest/schemas/well.schema
@@ -1,47 +1,59 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/0.4/schemas/well.schema",
- "title": "OME-NGFF well schema",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr well schema",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
- "well": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
"properties": {
- "images": {
- "description": "The fields of view for this well",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "acquisition": {
- "description": "A unique identifier within the context of the plate",
- "type": "integer"
- },
- "path": {
- "description": "The path for this field of view subgroup",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+$"
- }
- },
- "required": [
- "path"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
"version": {
- "description": "The version of the specification",
"type": "string",
"enum": [
"0.5-dev"
]
+ },
+ "well": {
+ "type": "object",
+ "properties": {
+ "images": {
+ "description": "The fields of view for this well",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "acquisition": {
+ "description": "A unique identifier within the context of the plate",
+ "type": "integer"
+ },
+ "path": {
+ "description": "The path for this field of view subgroup",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+$"
+ }
+ },
+ "required": [
+ "path"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ }
+ },
+ "required": [
+ "images"
+ ]
}
},
"required": [
- "images"
+ "version",
+ "well"
]
}
- }
+ },
+ "required": [
+ "ome"
+ ]
}
diff --git a/latest/transform-details.bs b/latest/transform-details.bs
new file mode 100644
index 00000000..f095b712
--- /dev/null
+++ b/latest/transform-details.bs
@@ -0,0 +1,107 @@
+
diff --git a/latest/schemas/bf2raw.schema b/latest/schemas/bf2raw.schema
index 834aee24..d661c8bf 100644
--- a/latest/schemas/bf2raw.schema
+++ b/latest/schemas/bf2raw.schema
@@ -1,14 +1,16 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/bf2raw.schema",
- "title": "NGFF container produced by bioformats2raw",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "Zarr container produced by bioformats2raw",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
"bioformats2raw.layout": {
"description": "The top-level identifier metadata added by bioformats2raw",
"type": "number",
- "enum": [3]
+ "enum": [
+ 3
+ ]
}
}
}
diff --git a/latest/schemas/image.schema b/latest/schemas/image.schema
index 3926c4f4..79f64f99 100644
--- a/latest/schemas/image.schema
+++ b/latest/schemas/image.schema
@@ -1,113 +1,136 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/image.schema",
- "title": "NGFF Image",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr Image",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
- "multiscales": {
- "description": "The multiscale datasets for this image",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "name": {
- "type": "string"
- },
- "datasets": {
- "type": "array",
- "minItems": 1,
- "items": {
- "type": "object",
- "properties": {
- "path": {
- "type": "string"
- },
- "coordinateTransformations": {
- "$ref": "#/$defs/coordinateTransformations"
- }
- },
- "required": ["path", "coordinateTransformations"]
- }
- },
- "version": {
- "type": "string",
- "enum": [
- "0.5-dev"
- ]
- },
- "axes": {
- "$ref": "#/$defs/axes"
- },
- "coordinateTransformations": {
- "$ref": "#/$defs/coordinateTransformations"
- }
- },
- "required": [
- "datasets", "axes"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "omero": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
"properties": {
- "channels": {
+ "version": {
+ "type": "string",
+ "enum": [
+ "0.5-dev"
+ ]
+ },
+ "multiscales": {
+ "description": "The multiscale datasets for this image",
"type": "array",
"items": {
"type": "object",
"properties": {
- "window": {
+ "name": {
+ "type": "string"
+ },
+ "datasets": {
+ "type": "array",
+ "minItems": 1,
+ "items": {
+ "type": "object",
+ "properties": {
+ "path": {
+ "type": "string"
+ },
+ "coordinateTransformations": {
+ "$ref": "#/$defs/coordinateTransformations"
+ }
+ },
+ "required": [
+ "path",
+ "coordinateTransformations"
+ ]
+ }
+ },
+ "axes": {
+ "$ref": "#/$defs/axes"
+ },
+ "coordinateTransformations": {
+ "$ref": "#/$defs/coordinateTransformations"
+ },
+ "labels": {
+ "type": "array",
+ "items": {
+ "type": "string"
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ }
+ },
+ "required": [
+ "datasets",
+ "axes"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "omero": {
+ "type": "object",
+ "properties": {
+ "channels": {
+ "type": "array",
+ "items": {
"type": "object",
"properties": {
- "end": {
- "type": "number"
+ "window": {
+ "type": "object",
+ "properties": {
+ "end": {
+ "type": "number"
+ },
+ "max": {
+ "type": "number"
+ },
+ "min": {
+ "type": "number"
+ },
+ "start": {
+ "type": "number"
+ }
+ },
+ "required": [
+ "start",
+ "min",
+ "end",
+ "max"
+ ]
+ },
+ "label": {
+ "type": "string"
},
- "max": {
- "type": "number"
+ "family": {
+ "type": "string"
},
- "min": {
- "type": "number"
+ "color": {
+ "type": "string"
},
- "start": {
- "type": "number"
+ "active": {
+ "type": "boolean"
}
},
"required": [
- "start",
- "min",
- "end",
- "max"
+ "window",
+ "color"
]
- },
- "label": {
- "type": "string"
- },
- "family": {
- "type": "string"
- },
- "color": {
- "type": "string"
- },
- "active": {
- "type": "boolean"
}
- },
- "required": [
- "window",
- "color"
- ]
- }
+ }
+ },
+ "required": [
+ "channels"
+ ]
}
},
"required": [
- "channels"
+ "multiscales",
+ "version"
]
- }
+ },
+ "required": [
+ "ome"
+ ]
},
- "required": [ "multiscales" ],
-
"$defs": {
"axes": {
"type": "array",
@@ -122,7 +145,9 @@
},
"type": {
"type": "string",
- "enum": ["space"]
+ "enum": [
+ "space"
+ ]
},
"units": {
"type": "string"
@@ -141,10 +166,17 @@
},
"type": {
"type": "string",
- "enum": ["channel", "time", "space"]
+ "enum": [
+ "channel",
+ "time",
+ "space"
+ ]
}
},
- "required": ["name", "type"]
+ "required": [
+ "name",
+ "type"
+ ]
},
{
"type": "object",
@@ -155,11 +187,17 @@
"type": {
"type": "string",
"not": {
- "enum": ["space", "time", "channel"]
+ "enum": [
+ "space",
+ "time",
+ "channel"
+ ]
}
}
},
- "required": ["name"]
+ "required": [
+ "name"
+ ]
}
]
}
@@ -205,7 +243,10 @@
}
}
},
- "required": ["type", "scale"]
+ "required": [
+ "type",
+ "scale"
+ ]
},
{
"type": "object",
@@ -224,7 +265,10 @@
}
}
},
- "required": ["type", "translation"]
+ "required": [
+ "type",
+ "translation"
+ ]
}
]
}
diff --git a/latest/schemas/label.schema b/latest/schemas/label.schema
index 8b23d577..ebca25c4 100644
--- a/latest/schemas/label.schema
+++ b/latest/schemas/label.schema
@@ -1,75 +1,87 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/label.schema",
- "title": "OME-NGFF labelled image schema",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr labelled image schema",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
+ "required": [
+ "ome"
+ ],
"properties": {
- "image-label": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
+ "required": [
+ "version"
+ ],
"properties": {
- "colors": {
- "description": "The colors for this label image",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "label-value": {
- "description": "The value of the label",
- "type": "number"
- },
- "rgba": {
- "description": "The RGBA color stored as an array of four integers between 0 and 255",
- "type": "array",
- "items": {
- "type": "integer",
- "minimum": 0,
- "maximum": 255
- },
- "minItems": 4,
- "maxItems": 4
- }
- },
- "required": [
- "label-value"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "properties": {
- "description": "The properties for this label image",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "label-value": {
- "description": "The pixel value for this label",
- "type": "integer"
- }
- },
- "required": [
- "label-value"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "source": {
- "description": "The source of this label image",
- "type": "object",
- "properties": {
- "image": {
- "type": "string"
- }
- }
- },
"version": {
"description": "The version of the specification",
"type": "string",
"enum": [
"0.5-dev"
]
+ },
+ "image-label": {
+ "type": "object",
+ "properties": {
+ "colors": {
+ "description": "The colors for this label image",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "label-value": {
+ "description": "The value of the label",
+ "type": "number"
+ },
+ "rgba": {
+ "description": "The RGBA color stored as an array of four integers between 0 and 255",
+ "type": "array",
+ "items": {
+ "type": "integer",
+ "minimum": 0,
+ "maximum": 255
+ },
+ "minItems": 4,
+ "maxItems": 4
+ }
+ },
+ "required": [
+ "label-value"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "properties": {
+ "description": "The properties for this label image",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "label-value": {
+ "description": "The pixel value for this label",
+ "type": "integer"
+ }
+ },
+ "required": [
+ "label-value"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "source": {
+ "description": "The source of this label image",
+ "type": "object",
+ "properties": {
+ "image": {
+ "type": "string"
+ }
+ }
+ }
+ }
}
}
}
diff --git a/latest/schemas/ome.schema b/latest/schemas/ome.schema
index bd600a2a..dd4872e8 100644
--- a/latest/schemas/ome.schema
+++ b/latest/schemas/ome.schema
@@ -1,8 +1,8 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/latest/schemas/ome.schema",
- "title": "NGFF group produced by bioformats2raw to contain OME metadata",
- "description": "JSON from OME-NGFF OME/.zattrs linked to an OME-XML file",
+ "title": "Zarr group produced by bioformats2raw to contain OME metadata",
+ "description": "JSON from OME-Zarr OME/zarr.json attributes linked to an OME-XML file",
"type": "object",
"properties": {
"series": {
diff --git a/latest/schemas/plate.schema b/latest/schemas/plate.schema
index 099f5bb4..150605e8 100644
--- a/latest/schemas/plate.schema
+++ b/latest/schemas/plate.schema
@@ -1,140 +1,156 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/0.4/schemas/plate.schema",
- "title": "OME-NGFF plate schema",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr plate schema",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
- "plate": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
"properties": {
- "acquisitions": {
- "description": "The acquisitions for this plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "id": {
- "description": "A unique identifier within the context of the plate",
- "type": "integer",
- "minimum": 0
- },
- "maximumfieldcount": {
- "description": "The maximum number of fields of view for the acquisition",
- "type": "integer",
- "exclusiveMinimum": 0
- },
- "name": {
- "description": "The name of the acquisition",
- "type": "string"
- },
- "description": {
- "description": "The description of the acquisition",
- "type": "string"
- },
- "starttime": {
- "description": "The start timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
- "type": "integer",
- "minimum": 0
- },
- "endtime": {
- "description": "The end timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
- "type": "integer",
- "minimum": 0
- }
- },
- "required": [
- "id"
- ]
- }
- },
"version": {
- "description": "The version of the specification",
"type": "string",
"enum": [
"0.5-dev"
]
},
- "field_count": {
- "description": "The maximum number of fields per view across all wells",
- "type": "integer",
- "exclusiveMinimum": 0
- },
- "name": {
- "description": "The name of the plate",
- "type": "string"
- },
- "columns": {
- "description": "The columns of the plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "name": {
- "description": "The column name",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+$"
+ "plate": {
+ "type": "object",
+ "properties": {
+ "acquisitions": {
+ "description": "The acquisitions for this plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "id": {
+ "description": "A unique identifier within the context of the plate",
+ "type": "integer",
+ "minimum": 0
+ },
+ "maximumfieldcount": {
+ "description": "The maximum number of fields of view for the acquisition",
+ "type": "integer",
+ "exclusiveMinimum": 0
+ },
+ "name": {
+ "description": "The name of the acquisition",
+ "type": "string"
+ },
+ "description": {
+ "description": "The description of the acquisition",
+ "type": "string"
+ },
+ "starttime": {
+ "description": "The start timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
+ "type": "integer",
+ "minimum": 0
+ },
+ "endtime": {
+ "description": "The end timestamp of the acquisition, expressed as epoch time i.e. the number seconds since the Epoch",
+ "type": "integer",
+ "minimum": 0
+ }
+ },
+ "required": [
+ "id"
+ ]
}
},
- "required": [
- "name"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "rows": {
- "description": "The rows of the plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "name": {
- "description": "The row name",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+$"
- }
+ "field_count": {
+ "description": "The maximum number of fields per view across all wells",
+ "type": "integer",
+ "exclusiveMinimum": 0
},
- "required": [
- "name"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
- "wells": {
- "description": "The wells of the plate",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "path": {
- "description": "The path to the well subgroup",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+/[A-Za-z0-9]+$"
+ "name": {
+ "description": "The name of the plate",
+ "type": "string"
+ },
+ "columns": {
+ "description": "The columns of the plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "name": {
+ "description": "The column name",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+$"
+ }
+ },
+ "required": [
+ "name"
+ ]
},
- "rowIndex": {
- "description": "The index of the well in the rows list",
- "type": "integer",
- "minimum": 0
+ "minItems": 1,
+ "uniqueItems": true
+ },
+ "rows": {
+ "description": "The rows of the plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "name": {
+ "description": "The row name",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+$"
+ }
+ },
+ "required": [
+ "name"
+ ]
},
- "columnIndex": {
- "description": "The index of the well in the columns list",
- "type": "integer",
- "minimum": 0
- }
+ "minItems": 1,
+ "uniqueItems": true
},
- "required": [
- "path", "rowIndex", "columnIndex"
- ]
+ "wells": {
+ "description": "The wells of the plate",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "path": {
+ "description": "The path to the well subgroup",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+/[A-Za-z0-9]+$"
+ },
+ "rowIndex": {
+ "description": "The index of the well in the rows list",
+ "type": "integer",
+ "minimum": 0
+ },
+ "columnIndex": {
+ "description": "The index of the well in the columns list",
+ "type": "integer",
+ "minimum": 0
+ }
+ },
+ "required": [
+ "path",
+ "rowIndex",
+ "columnIndex"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ }
},
- "minItems": 1,
- "uniqueItems": true
+ "required": [
+ "columns",
+ "rows",
+ "wells"
+ ]
}
},
"required": [
- "columns", "rows", "wells"
+ "version",
+ "plate"
]
}
- }
+ },
+ "required": [
+ "ome"
+ ]
}
diff --git a/latest/schemas/strict_image.schema b/latest/schemas/strict_image.schema
index bcecc003..3499be75 100644
--- a/latest/schemas/strict_image.schema
+++ b/latest/schemas/strict_image.schema
@@ -9,11 +9,13 @@
"multiscales": {
"items": {
"required": [
- "version", "metadata", "type", "name"
+ "metadata",
+ "type",
+ "name"
]
}
}
}
}
]
-}
\ No newline at end of file
+}
diff --git a/latest/schemas/strict_label.schema b/latest/schemas/strict_label.schema
index c52dd51b..9d888276 100644
--- a/latest/schemas/strict_label.schema
+++ b/latest/schemas/strict_label.schema
@@ -8,7 +8,6 @@
"properties": {
"image-label": {
"required": [
- "version",
"colors"
]
}
diff --git a/latest/schemas/strict_plate.schema b/latest/schemas/strict_plate.schema
index 5a88ab72..4c68707f 100644
--- a/latest/schemas/strict_plate.schema
+++ b/latest/schemas/strict_plate.schema
@@ -18,8 +18,7 @@
}
},
"required": [
- "name",
- "version"
+ "name"
]
}
}
diff --git a/latest/schemas/strict_well.schema b/latest/schemas/strict_well.schema
index 1e200294..e70dff9f 100644
--- a/latest/schemas/strict_well.schema
+++ b/latest/schemas/strict_well.schema
@@ -7,9 +7,7 @@
{
"properties": {
"well": {
- "required": [
- "version"
- ]
+ "required": []
}
}
}
diff --git a/latest/schemas/well.schema b/latest/schemas/well.schema
index 891abd05..651b8dc0 100644
--- a/latest/schemas/well.schema
+++ b/latest/schemas/well.schema
@@ -1,47 +1,59 @@
{
"$schema": "https://json-schema.org/draft/2020-12/schema",
"$id": "https://ngff.openmicroscopy.org/0.4/schemas/well.schema",
- "title": "OME-NGFF well schema",
- "description": "JSON from OME-NGFF .zattrs",
+ "title": "OME-Zarr well schema",
+ "description": "JSON from OME-Zarr zarr.json attributes",
"type": "object",
"properties": {
- "well": {
+ "ome": {
+ "description": "The OME namespace for metadata",
"type": "object",
"properties": {
- "images": {
- "description": "The fields of view for this well",
- "type": "array",
- "items": {
- "type": "object",
- "properties": {
- "acquisition": {
- "description": "A unique identifier within the context of the plate",
- "type": "integer"
- },
- "path": {
- "description": "The path for this field of view subgroup",
- "type": "string",
- "pattern": "^[A-Za-z0-9]+$"
- }
- },
- "required": [
- "path"
- ]
- },
- "minItems": 1,
- "uniqueItems": true
- },
"version": {
- "description": "The version of the specification",
"type": "string",
"enum": [
"0.5-dev"
]
+ },
+ "well": {
+ "type": "object",
+ "properties": {
+ "images": {
+ "description": "The fields of view for this well",
+ "type": "array",
+ "items": {
+ "type": "object",
+ "properties": {
+ "acquisition": {
+ "description": "A unique identifier within the context of the plate",
+ "type": "integer"
+ },
+ "path": {
+ "description": "The path for this field of view subgroup",
+ "type": "string",
+ "pattern": "^[A-Za-z0-9]+$"
+ }
+ },
+ "required": [
+ "path"
+ ]
+ },
+ "minItems": 1,
+ "uniqueItems": true
+ }
+ },
+ "required": [
+ "images"
+ ]
}
},
"required": [
- "images"
+ "version",
+ "well"
]
}
- }
+ },
+ "required": [
+ "ome"
+ ]
}
diff --git a/latest/transform-details.bs b/latest/transform-details.bs
new file mode 100644
index 00000000..f095b712
--- /dev/null
+++ b/latest/transform-details.bs
@@ -0,0 +1,107 @@
++Title: Coordinates and Transformations +Shortname: ome-ngff-transformations +Level: 1 +Status: LS-COMMIT +Status: w3c/ED +Group: ome +URL: https://ngff.openmicroscopy.org/latest/ +Repository: https://github.com/ome/ngff +Issue Tracking: Forums https://forum.image.sc/tag/ome-ngff +Logo: http://www.openmicroscopy.org/img/logos/ome-logomark.svg +Local Boilerplate: header no +Local Boilerplate: copyright no +Boilerplate: style-darkmode off +Markup Shorthands: markdown yes +Editor: Josh Moore, Open Microscopy Environment (OME) https://www.openmicroscopy.org +Editor: Sébastien Besson, Open Microscopy Environment (OME) https://www.openmicroscopy.org +Editor: Constantin Pape, European Molecular Biology Laboratory (EMBL) https://www.embl.org/sites/heidelberg/ +Editor: John Bogovic, Hughes Medical Institute Janelia (HHMI) https://www.janelia.org/ +Editor: Norman Rzepka, scalable minds https://scalableminds.com/, https://orcid.org/0000-0002-8168-7929 +Abstract: This document contains next-generation file format (NGFF) +Abstract: specifications for storing bioimaging data in the cloud. +Abstract: All specifications are submitted to the https://image.sc community for review. +Status Text: The current released version of this specification is +Status Text: will be provided between numbered versions. Data written with these latest changes +Status Text: (an "editor's draft") will not necessarily be supported. ++ +Coordinates and Axes {#coords-axes} +===================== + +OME-Zarr datasets are arrays that hold values. The arrays may be indexed by discrete (integer) +coordinates in order to obtain a corresponding value. If values are desired at continuous (real-valued) +coordinates, then interpolation is required. + +Interpolation {#interp} +--------------------- + +Interpolation is the process of producing values at continuous coordinates from data sampled at discrete +coordinates. "Nearest-neighbor" and "N-Linear" are the two most commonly used interpolation methods. + + +Pixel coordinates {#pix-coords} +--------------------- + +**The pixel center is the origin of the continuous coordinate system.** + +### Top-left convention + +A common alternative convention is for the origin in the continuous space is at the "top-left" of the pixel. +This is not recommended, but can be acheived by explicitly adding a half-pixel translation, for example: + +```json +{ + "name": "center_to_top-left", + "type": "translation", + "translation" : [0.5, 0.5], + "output_space" : "top-left-space" +} +``` + +Coordinate Transformations {#coord-tforms} +===================== + +This document describes background and motivation that is outside the NGFF specification. + + +Direction {#direction} +--------------------- + +Specified coordinate transforms are in the "forward" direction. They represent functions +from *points* in the input space to *points* in the output space. For example, the transformation `"ij2xy"` + + +```json +{ + "name": "ij2xy", + "type": "scale", + "scale": [2, 0.5] + "input_axes" : ["i", "j"] + "output_axes" : ["x", "y"] +} +``` + +representes the function + +``` +x = 2 * i +y = 0.5 * j +``` + + +Recommendations {#recommendations} +===================== + + +"Native" physical space +--------------------- + +Datasets SHOULD define a transformation from array space to their "native physical space." +This transformation SHOULD describe physical pixel spacing and origin only, and therefore SHOULD consist of +`scale` and/or `translation` types only. + +Subsequent reorientation / registration transformations SHOULD use this native space as their `input_space`, +i.e., transformations should be defined in physical coordinates. + +