From a9c7e3ecb73c48bb27a46d3e68fb591d2824aae8 Mon Sep 17 00:00:00 2001
From: xusuyong <2209245477@qq.com>
Date: Wed, 22 Nov 2023 20:26:08 +0800
Subject: [PATCH 1/2] add RegAE example
---
docs/zh/examples/RegAE.md | 357 +++++++++++++++++++++++++++++++++
examples/RegAE/RegAE.py | 177 ++++++++++++++++
examples/RegAE/conf/RegAE.yaml | 51 +++++
examples/RegAE/dataloader.py | 142 +++++++++++++
ppsci/arch/__init__.py | 2 +
ppsci/arch/vae.py | 74 +++++++
ppsci/loss/__init__.py | 2 +
ppsci/loss/kl.py | 45 +++++
8 files changed, 850 insertions(+)
create mode 100644 docs/zh/examples/RegAE.md
create mode 100644 examples/RegAE/RegAE.py
create mode 100644 examples/RegAE/conf/RegAE.yaml
create mode 100644 examples/RegAE/dataloader.py
create mode 100644 ppsci/arch/vae.py
create mode 100644 ppsci/loss/kl.py
diff --git a/docs/zh/examples/RegAE.md b/docs/zh/examples/RegAE.md
new file mode 100644
index 000000000..bbd874c0f
--- /dev/null
+++ b/docs/zh/examples/RegAE.md
@@ -0,0 +1,357 @@
+# Learning to regularize with a variational autoencoder for hydrologic inverse analysis
+
+## 1.简介
+
+本项目基于paddle框架复现。
+
+论文主要点如下:
+* 作者提出了一种基于变分自动编码器 (VAE)的正则化方法;
+* 这种方法的优点1: 对来自VAE的潜在变量(此处指encoder的输出)执行正则化,可以简单地对其进行正则化;
+* 这种方法的优点2: VAE减少了优化问题中的变量数量,从而在伴随方法不可用时使基于梯度的优化在计算上更加高效。
+
+本项目关键技术要点:
+
+* 实现paddle和julia混写代码梯度传递,避免大面积重写julia代码并能够调用优秀的julia代码;
+* 发现paddle minimize_lbfgs存在问题, 待提交issue确认。
+
+
+实验结果要点:
+* 成功复现论文代码框架及全流程运行测试;
+* 本次复现精度因无法使用相同样本,无法与论文中数据进行一一比较。本项目给出了采用paddle编写的框架结果。
+
+论文信息:
+O'Malley D, Golden J K, Vesselinov V V. Learning to regularize with a variational autoencoder for hydrologic inverse analysis[J]. arXiv preprint arXiv:1906.02401, 2019.
+
+参考GitHub地址:
+https://github.com/madsjulia/RegAE.jl
+
+项目aistudio地址:
+https://aistudio.baidu.com/aistudio/projectdetail/5541961
+
+模型结构
+
+
+
+
+
+## 2.数据集
+
+本项目数据集通过作者提供的julia代码生成,生成后保存为npz文件,已上传aistudio[数据集](https://aistudio.baidu.com/aistudio/datasetdetail/193595)并关联本项目。
+
+以下为数据生成和数据保存代码的说明
+
+(1)作者通过julia中的DPFEHM和GaussianRandomFields进行数据生成,代码可参考本项目/home/aistudio/RegAE.jl/examples/hydrology/ex_gaussian.jl,可根据其中参数进行修改;
+
+(2)数据保存代码。在/home/aistudio/RegAE.jl/examples/hydrology/ex.jl代码中增加以下代码,可将数据通过转换为numpy数据并保存为npz。
+
+```julia
+using Distributed
+using PyCall # 增加此处引用
+
+@everywhere variablename = "allloghycos"
+@everywhere datafilename = "$(results_dir)/trainingdata.jld2"
+if !isfile(datafilename)
+ if nworkers() == 1
+ error("Please run in parallel: julia -p 32")
+ end
+ numsamples = 10^5
+ @time allloghycos = SharedArrays.SharedArray{Float32}(numsamples, ns[2], ns[1]; init=A -> samplehyco!(A; setseed=true))
+ # @time JLD2.@save datafilename allloghycos
+
+ ########### 此处为增加部分 ###########
+ p_trues = SharedArrays.SharedArray{Float32}(3, ns[2], ns[1]; init=samplehyco!) # 计算p_true
+
+ np = pyimport("numpy")
+ training_data = np.asarray(allloghycos)
+ test_data = np.asarray(p_trues)
+
+ np_coords = np.asarray(coords)
+ np_neighbors = np.asarray(neighbors)
+ np_areasoverlengths = np.asarray(areasoverlengths)
+ np_dirichletnodes = np.asarray(dirichletnodes)
+ np_dirichletheads = np.asarray(dirichletheads)
+
+ np.savez("$(results_dir)/gaussian_train.npz",
+ data=training_data,
+ test_data=test_data,
+ coords=np_coords,
+ neighbors=np_neighbors,
+ areasoverlengths=np_areasoverlengths,
+ dirichletnodes=np_dirichletnodes,
+ dirichletheads=np_dirichletheads)
+end
+```
+
+## 数据标准化
+
+* 数据标准化方式: $z = (x - \mu)/ \sigma$
+
+
+```python
+ class ScalerStd(object):
+ """
+ Desc: Normalization utilities with std mean
+ """
+
+ def __init__(self):
+ self.mean = 0.
+ self.std = 1.
+
+ def fit(self, data):
+ self.mean = np.mean(data)
+ self.std = np.std(data)
+
+ def transform(self, data):
+ mean = paddle.to_tensor(self.mean).type_as(data).to(
+ data.device) if paddle.is_tensor(data) else self.mean
+ std = paddle.to_tensor(self.std).type_as(data).to(
+ data.device) if paddle.is_tensor(data) else self.std
+ return (data - mean) / std
+
+ def inverse_transform(self, data):
+ mean = paddle.to_tensor(self.mean) if paddle.is_tensor(data) else self.mean
+ std = paddle.to_tensor(self.std) if paddle.is_tensor(data) else self.std
+ return (data * std) + mean
+```
+
+## 定义Dataset
+
+1. 通过读取预保存npz加载数据,当前数据类型为 [data_nums, 100, 100], 此处100为数据生成过程中指定
+2. 数据reshape为 [data_nums, 10000]
+3. 数据划分为8:2用与train和test
+4. 通过对train数据得到标准化参数mean和std,并用此参数标准化train和test数据集
+5. 通过dataloader得到的数据形式为 [batch_size, 10000]
+```python
+ class CustomDataset(Dataset):
+ def __init__(self, file_path, data_type="train"):
+ """
+
+ :param file_path:
+ :param data_type: train or test
+ """
+ super().__init__()
+ all_data = np.load(file_path)
+ data = all_data["data"]
+ num, _, _ = data.shape
+ data = data.reshape(num, -1)
+
+ self.neighbors = all_data['neighbors']
+ self.areasoverlengths = all_data['areasoverlengths']
+ self.dirichletnodes = all_data['dirichletnodes']
+ self.dirichleths = all_data['dirichletheads']
+ self.Qs = np.zeros([all_data['coords'].shape[-1]])
+ self.val_data = all_data["test_data"]
+
+ self.data_type = data_type
+
+ self.train_len = int(num * 0.8)
+ self.test_len = num - self.train_len
+
+ self.train_data = data[:self.train_len]
+ self.test_data = data[self.train_len:]
+
+ self.scaler = ScalerStd()
+ self.scaler.fit(self.train_data)
+
+ self.train_data = self.scaler.transform(self.train_data)
+ self.test_data = self.scaler.transform(self.test_data)
+
+ def __getitem__(self, idx):
+ if self.data_type == "train":
+ return self.train_data[idx]
+ else:
+ return self.test_data[idx]
+
+ def __len__(self):
+ if self.data_type == "train":
+ return self.train_len
+ else:
+ return self.test_len
+```
+## 将数据转换为IterableNPZDataset的形式
+
+```python
+np.savez("data.npz", p_train=train_data.train_data, p_test=train_data.test_data)
+```
+
+## 3.环境依赖
+
+本项目为julia和python混合项目。
+
+### julia依赖
+
+* DPFEHM
+* Zygote
+
+### python依赖
+* paddle
+* julia (pip安装)
+* matplotlib
+
+本项目已经提供安装后压缩文档,可fork本项目后执行以下代码进行解压安装。
+
+
+```python
+# 解压预下载文件和预编译文件
+!tar zxf /home/aistudio/opt/curl-7.88.1.tar.gz -C /home/aistudio/opt # curl 预下载文件
+!tar zxf /home/aistudio/opt/curl-7.88.1-build.tgz -C /home/aistudio/opt # curl 预编译文件
+!tar zxf /home/aistudio/opt/julia-1.8.5-linux-x86_64.tar.gz -C /home/aistudio/opt # julia 预下载文件
+!tar zxf /home/aistudio/opt/julia_package.tgz -C /home/aistudio/opt # julia依赖库文件
+!tar zxf /home/aistudio/opt/external-libraries.tgz -C /home/aistudio/opt # pip依赖库文件
+```
+
+
+```python
+####### 以下指令需要时可参考执行,上述压缩包已经完成以下内容 #######
+
+# curl 编译指令,当解压后无效使用
+!mkdir -p /home/aistudio/opt/curl-7.88.1-build
+!/home/aistudio/opt/curl-7.88.1/configure --prefix=/home/aistudio/opt/curl-7.88.1-build --with-ssl --enable-tls-srp
+!make install -j4
+
+# 指定curl预编译文件
+!export LD_LIBRARY_PATH=/home/aistudio/opt/curl-7.88.1-build/lib:$LD_LIBRARY_PATH
+!export PATH=/home/aistudio/opt/curl-7.88.1-build/bin:$PATH
+!export CPATH=/home/aistudio/opt/curl-7.88.1-build/include:$CPATH
+!export LIBRARY_PATH=/home/aistudio/opt/curl-7.88.1-build/lib:$LIBRARY_PATH
+
+# 指定已经安装的julia包
+!export JULIA_DEPOT_PATH=/home/aistudio/opt/julia_package
+# 指定julia使用清华源
+!export JULIA_PKG_SERVER=https://mirrors.tuna.tsinghua.edu.cn/julia
+# julia 安装依赖库
+# 需要先export JULIA_DEPOT_PATH 环境变量,否则安装位置为~/.julia, aistudio无法保存
+!/home/aistudio/opt/julia-1.8.5/bin/julia -e "using Pkg; Pkg.add(\"DPFEHM\")"
+!/home/aistudio/opt/julia-1.8.5/bin/julia -e "using Pkg; Pkg.add(\"Zygote\")"
+!/home/aistudio/opt/julia-1.8.5/bin/julia -e "using Pkg; Pkg.add(\"PyCall\")"
+```
+
+使用方法可以参考以下代码和julia导数传递测试.ipynb文件。
+
+```python
+import paddle
+import os
+import sys
+
+# julia 依赖
+os.environ['JULIA_DEPOT_PATH'] = '/home/aistudio/opt/julia_package'
+# pip 依赖
+sys.path.append('/home/aistudio/opt/external-libraries')
+
+# julieries
+from julia.api import Julia
+
+jl = Julia(compiled_modules=False,runtime="/home/aistudio/opt/julia-1.8.5/bin/julia")
+# import julia
+from julia import Main
+```
+
+## 4.快速开始
+
+本项目运行分为两个步骤:
+* (1)训练步骤。通过运行train.ipynb文件,可以得到训练后的模型参数,具体代码请参考train.ipynb文件及其中注释说明;
+* (2)测试步骤。通过运行test.ipynb文件,应用训练后的模型参数,对latent domain进行优化。
+
+## 5.代码结构与详细说明
+```text
+├── data #预生成数据文件
+│ └── data193595
+├── main.ipynb #本说明文件
+├── opt #环境配置文件,已压缩,解压即可使用
+│ ├── curl-7.88.1
+│ ├── curl-7.88.1-build
+│ ├── curl-7.88.1-build.tgz
+│ ├── curl-7.88.1.tar.gz
+│ ├── external-libraries
+│ ├── external-libraries.tgz
+│ ├── julia-1.8.5
+│ ├── julia-1.8.5-linux-x86_64.tar.gz
+│ ├── julia_package
+│ └── julia_package.tgz
+├── params_vae_nz100 #模型参数文件
+│ └── model.pdparams
+├── params_vae_nz200
+│ └── model.pdparams
+├── params_vae_nz400
+│ └── model.pdparams
+├── test.ipynb #测试文件
+├── train.ipynb #训练文件
+├── julia导数传递测试.ipynb #julia和python混合测试文件
+```
+
+### train文件和test文件关联性说明
+
+我们依照论文作者的符号进行说明,$p$为数据输入,$\hat{p}$为数据输出,$loss=mse(p,\hat{p}) + loss_{kl}(\hat{p},N(0,1))$。
+
+* (1)通过train能够得到训练后的Autoencoder(包含encoder和decoder);
+* (2)通过test调用训练后的encoder针对testdata生成latent_test,并得到latent_mean;
+* (3)针对新生成的样本$p_{new}$,通过LBFGS方法不断优化latent_mean,直到obj_fun最小,其中obj_fun = mse($p_{new}$,$\hat{p}_{new}$)+mse(sci_fun($p_{new}$),sci_fun($\hat{p}_{new}$)),sci_fun为任何其他科学计算模拟方法。
+
+### paddle.incubate.optimizer.functional.minimize_lbfgs 问题
+
+以下为paddle官方minimize_lbfgs API:
+```python
+paddle.incubate.optimizer.functional.minimize_lbfgs(objective_func, initial_position, history_size=100, max_iters=50, tolerance_grad=1e-08, tolerance_change=1e-08, initial_inverse_hessian_estimate=None, line_search_fn='strong_wolfe', max_line_search_iters=50, initial_step_length=1.0, dtype='float32', name=None)
+```
+
+
+* (1)参数max_line_search_iters无效。虽然设置了此参数,但是内部没有传递对应参数;
+* (2)中wolfe条件1错误。line256处应为`phi_2 >= phi_1`,以下为paddle部分源码。
+
+```python
+ # 1. If phi(a2) > phi(0) + c_1 * a2 * phi'(0) or [phi(a2) >= phi(a1) and i > 1],
+ # a_star= zoom(a1, a2) and stop;
+ pred1 = ~done & ((phi_2 > phi_0 + c1 * a2 * derphi_0) |
+ ((phi_2 >= phi_0) & (i > 1)))
+```
+
+## 6.复现结果
+
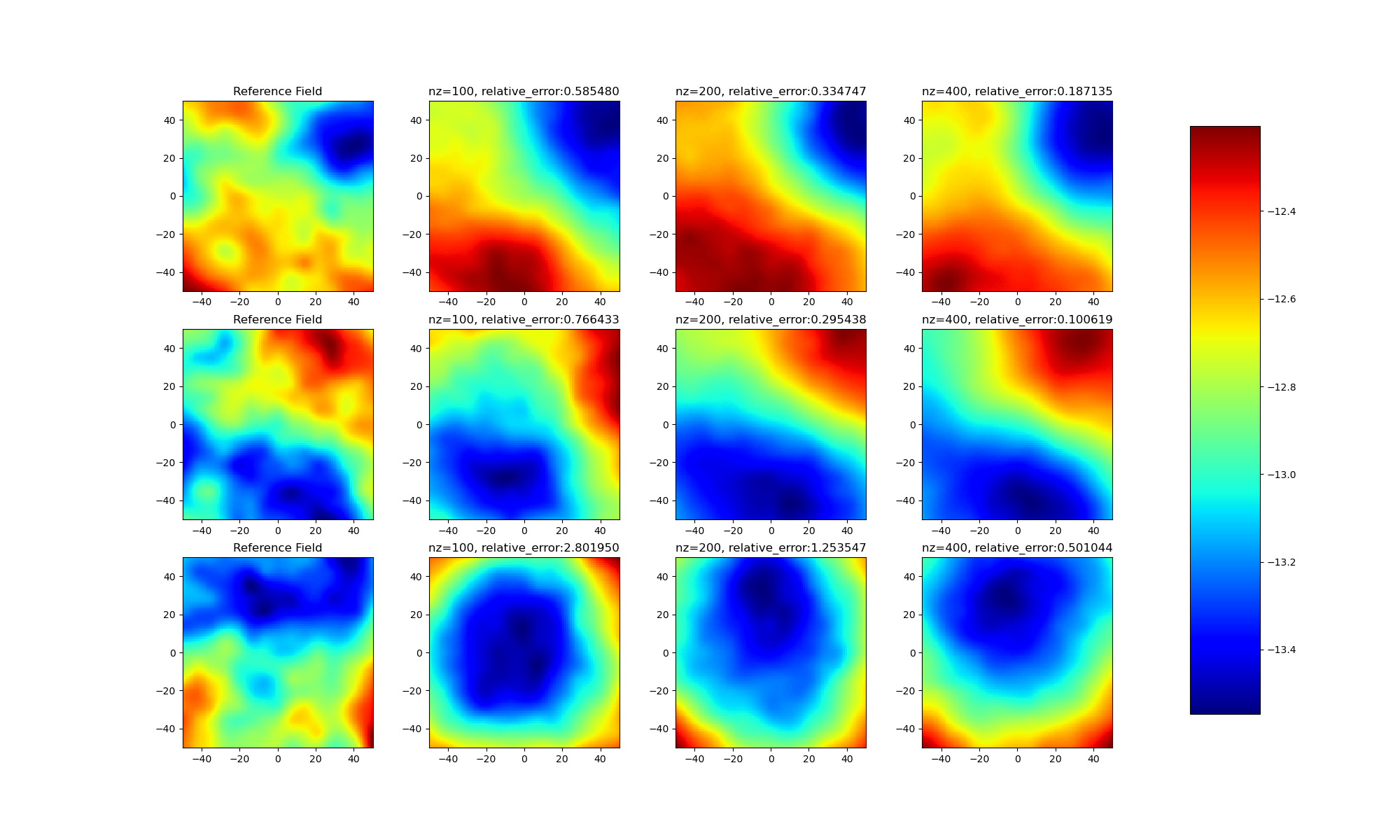
+### 不同latent维度对比
+
+
+通过实验结果可以发现:
+* (1)不同样本之间存在差距,并不是所有样本都能优化得到良好的latent变量;
+* (2)随着模型latent维度的上升,模型效果逐渐提升。
+
+### latent_random和latent_mean对比
+本项目还增加了latent_random和latent_mean对生成结果的对比。此处对latent_random和latent_mean再次说明:
+* latent_random:通过paddle.randn生成的高斯噪声得到;
+* latent_mean:通过对所有testdata进行encoder结果平均得到。
+
+以下为通过latent_random得到的实验结果
+
+
+通过对比,可以发现latent_mean对优化结果重要影响。近似正确的latent变量能够得到更优的生成结果。
+
+### LBFGS优化收敛情况
+可以从如下图中看出,使用paddle minimize_lbfgs能够有效优化收敛。
+
+
+
+
+
+## 7.延伸思考
+
+如果深入思考本项目,会发现模型在test过程中是使用真实数据作为目标进行lbfgs优化,这种计算方式还有意义吗?
+
+回答是肯定的!有意义!
+
+以下为本人个人观点:
+* (1)通过实验对比latent_random和latent_mean的最终生成结果差距,可以发现一个良好的初值对模型的影响是巨大的。当前diffusion模型在sample生成过程对生成的高斯噪声不断做denoise操作,这其中生成的噪声数据如果经过预先优化,不仅能够加速diffusion的生成速度,而且能够提升数据的生成质量。
+* (2)在域迁移等研究领域,可以使用这种latent逐渐生成中间过渡变量,达到不同域数据的迁移生成。
+
+
+
+## 7.模型信息
+
+| 信息 | 说明|
+| -------- | -------- |
+| 发布者 | 朱卫国 (DrownFish19) |
+| 发布时间 | 2023.03 |
+| 框架版本 | paddle 2.4.1 |
+| 支持硬件 | GPU、CPU |
+| aistudio | [notebook](https://aistudio.baidu.com/aistudio/projectdetail/5541961) |
+
+请点击[此处](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576)查看本环境基本用法.
+Please click [here ](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576) for more detailed instructions.
diff --git a/examples/RegAE/RegAE.py b/examples/RegAE/RegAE.py
new file mode 100644
index 000000000..3170429a9
--- /dev/null
+++ b/examples/RegAE/RegAE.py
@@ -0,0 +1,177 @@
+# Copyright (c) 2023 PaddlePaddle Authors. All Rights Reserved.
+
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in compliance with the License.
+# You may obtain a copy of the License at
+
+# http://www.apache.org/licenses/LICENSE-2.0
+
+# Unless required by applicable law or agreed to in writing, software
+# distributed under the License is distributed on an "AS IS" BASIS,
+# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
+# See the License for the specific language governing permissions and
+# limitations under the License.
+
+from __future__ import annotations
+
+from os import path as osp
+
+import hydra
+from omegaconf import DictConfig
+from paddle import nn
+
+import ppsci
+from ppsci.loss import KLLoss
+from ppsci.utils import logger
+
+criterion = nn.MSELoss()
+kl_loss = KLLoss()
+
+
+def loss_expr(output_dict, label_dict, weight_dict=None):
+
+ return kl_loss(output_dict) + criterion(output_dict["p_hat"], label_dict["p_hat"])
+
+
+def train(cfg: DictConfig):
+ # set random seed for reproducibility
+ ppsci.utils.misc.set_random_seed(cfg.seed)
+ # initialize logger
+ logger.init_logger("ppsci", osp.join(cfg.output_dir, "train.log"), "info")
+
+ # set model
+ model = ppsci.arch.AutoEncoder(**cfg.MODEL)
+
+ # set dataloader config
+ train_dataloader_cfg = {
+ "dataset": {
+ "name": "IterableNPZDataset",
+ "file_path": cfg.TRAIN_FILE_PATH,
+ "input_keys": ("p",),
+ "label_keys": ("p_hat",),
+ "alias_dict": {"p": "p_train", "p_hat": "p_train"},
+ },
+ "batch_size": cfg.TRAIN.batch_size,
+ "sampler": {
+ "name": "BatchSampler",
+ "drop_last": False,
+ "shuffle": True,
+ },
+ }
+
+ # set constraint
+ sup_constraint = ppsci.constraint.SupervisedConstraint(
+ train_dataloader_cfg,
+ output_expr={"p_hat": lambda out: out["p_hat"]},
+ loss=ppsci.loss.FunctionalLoss(loss_expr),
+ name="Sup",
+ )
+ # wrap constraints together
+ constraint = {sup_constraint.name: sup_constraint}
+
+ # set optimizer
+ optimizer = ppsci.optimizer.Adam(cfg.TRAIN.learning_rate)(model)
+
+ # set validator
+ eval_dataloader_cfg = {
+ "dataset": {
+ "name": "IterableNPZDataset",
+ "file_path": cfg.VALID_FILE_PATH,
+ "input_keys": ("p",),
+ "label_keys": ("p_hat",),
+ "alias_dict": {"p": "p_train", "p_hat": "p_train"},
+ },
+ "batch_size": cfg.EVAL.batch_size,
+ "sampler": {
+ "name": "BatchSampler",
+ "drop_last": False,
+ "shuffle": False,
+ },
+ }
+ sup_validator = ppsci.validate.SupervisedValidator(
+ eval_dataloader_cfg,
+ loss=ppsci.loss.MSELoss(),
+ output_expr={"p_hat": lambda out: out["p_hat"]},
+ metric={"L2Rel": ppsci.metric.L2Rel()},
+ )
+ validator = {sup_validator.name: sup_validator}
+
+ # initialize solver
+ solver = ppsci.solver.Solver(
+ model,
+ constraint,
+ cfg.output_dir,
+ optimizer,
+ None,
+ cfg.TRAIN.epochs,
+ cfg.TRAIN.iters_per_epoch,
+ save_freq=cfg.TRAIN.save_freq,
+ eval_during_train=cfg.TRAIN.eval_during_train,
+ eval_freq=cfg.TRAIN.eval_freq,
+ validator=validator,
+ eval_with_no_grad=cfg.EVAL.eval_with_no_grad,
+ )
+ # train model
+ solver.train()
+ # evaluate after finished training
+ solver.eval()
+
+
+def evaluate(cfg: DictConfig):
+ # set random seed for reproducibility
+ ppsci.utils.misc.set_random_seed(cfg.seed)
+ # initialize logger
+ logger.init_logger("ppsci", osp.join(cfg.output_dir, "train.log"), "info")
+
+ # set model
+ model = ppsci.arch.AutoEncoder(**cfg.MODEL)
+
+ # set validator
+ eval_dataloader_cfg = {
+ "dataset": {
+ "name": "IterableNPZDataset",
+ "file_path": cfg.VALID_FILE_PATH,
+ "input_keys": ("p",),
+ "label_keys": ("p_hat",),
+ "alias_dict": {"p": "p_train", "p_hat": "p_train"},
+ },
+ "batch_size": cfg.EVAL.batch_size,
+ "sampler": {
+ "name": "BatchSampler",
+ "drop_last": False,
+ "shuffle": False,
+ },
+ }
+ sup_validator = ppsci.validate.SupervisedValidator(
+ eval_dataloader_cfg,
+ loss=ppsci.loss.MSELoss(),
+ output_expr={"p_hat": lambda out: out["p_hat"]},
+ metric={"L2Rel": ppsci.metric.L2Rel()},
+ )
+ validator = {sup_validator.name: sup_validator}
+
+ # initialize solver
+ solver = ppsci.solver.Solver(
+ model,
+ None,
+ output_dir=cfg.output_dir,
+ validator=validator,
+ pretrained_model_path=cfg.EVAL.pretrained_model_path,
+ eval_with_no_grad=cfg.EVAL.eval_with_no_grad,
+ )
+ # evaluate after finished training
+ solver.eval()
+
+
+@hydra.main(version_base=None, config_path="./conf", config_name="RegAE.yaml")
+def main(cfg: DictConfig):
+ if cfg.mode == "train":
+ train(cfg)
+ elif cfg.mode == "eval":
+ evaluate(cfg)
+ else:
+ raise ValueError(f"cfg.mode should in ['train', 'eval'], but got '{cfg.mode}'")
+
+
+if __name__ == "__main__":
+ main()
diff --git a/examples/RegAE/conf/RegAE.yaml b/examples/RegAE/conf/RegAE.yaml
new file mode 100644
index 000000000..133ac82a5
--- /dev/null
+++ b/examples/RegAE/conf/RegAE.yaml
@@ -0,0 +1,51 @@
+hydra:
+ run:
+ # dynamic output directory according to running time and override name
+ dir: output_RegAE/${now:%Y-%m-%d}/${now:%H-%M-%S}/${hydra.job.override_dirname}
+ job:
+ name: ${mode} # name of logfile
+ chdir: false # keep current working directory unchanged
+ config:
+ override_dirname:
+ exclude_keys:
+ - TRAIN.checkpoint_path
+ - TRAIN.pretrained_model_path
+ - EVAL.pretrained_model_path
+ - mode
+ - output_dir
+ - log_freq
+ sweep:
+ # output directory for multirun
+ dir: ${hydra.run.dir}
+ subdir: ./
+
+# general settings
+mode: train # running mode: train/eval
+seed: 42
+output_dir: ${hydra:run.dir}
+TRAIN_FILE_PATH: data.npz
+VALID_FILE_PATH: data.npz
+
+# model settings
+MODEL:
+ input_keys: [ "p",]
+ output_keys: ["mu", "log_sigma", "p_hat"]
+ input_dim: 10000
+ latent_dim: 100
+ hidden_dim: 100
+
+# training settings
+TRAIN:
+ epochs: 200
+ iters_per_epoch: 1
+ eval_during_train: true
+ save_freq: 200
+ eval_freq: 200
+ learning_rate: 0.0001
+ batch_size: 128
+
+# evaluation settings
+EVAL:
+ pretrained_model_path: null
+ eval_with_no_grad: False
+ batch_size: 128
diff --git a/examples/RegAE/dataloader.py b/examples/RegAE/dataloader.py
new file mode 100644
index 000000000..e64654b96
--- /dev/null
+++ b/examples/RegAE/dataloader.py
@@ -0,0 +1,142 @@
+"""
+输入数据类型 10^5 * 100 * 100
+
+1.按照8:2划分训练数据集和测试数据集
+2.通过训练数据进行标准正则化
+"""
+import numpy as np
+import paddle
+from paddle.io import DataLoader
+from paddle.io import Dataset
+
+
+class ScalerStd(object):
+ """
+ Desc: Normalization utilities with std mean
+ """
+
+ def __init__(self):
+ self.mean = 0.0
+ self.std = 1.0
+
+ def fit(self, data):
+ self.mean = np.mean(data)
+ self.std = np.std(data)
+
+ def transform(self, data):
+ mean = (
+ paddle.to_tensor(self.mean).type_as(data).to(data.device)
+ if paddle.is_tensor(data)
+ else self.mean
+ )
+ std = (
+ paddle.to_tensor(self.std).type_as(data).to(data.device)
+ if paddle.is_tensor(data)
+ else self.std
+ )
+ return (data - mean) / std
+
+ def inverse_transform(self, data):
+ mean = paddle.to_tensor(self.mean) if paddle.is_tensor(data) else self.mean
+ std = paddle.to_tensor(self.std) if paddle.is_tensor(data) else self.std
+ return (data * std) + mean
+
+
+class ScalerMinMax(object):
+ """
+ Desc: Normalization utilities with min max
+ """
+
+ def __init__(self):
+ self.min = 0.0
+ self.max = 1.0
+
+ def fit(self, data):
+ self.min = np.min(data, axis=0)
+ self.max = np.max(data, axis=0)
+
+ def transform(self, data):
+ _min = (
+ paddle.to_tensor(self.min).type_as(data).to(data.device)
+ if paddle.is_tensor(data)
+ else self.min
+ )
+ _max = (
+ paddle.to_tensor(self.max).type_as(data).to(data.device)
+ if paddle.is_tensor(data)
+ else self.max
+ )
+ data = 1.0 * (data - _min) / (_max - _min)
+ return 2.0 * data - 1.0
+
+ def inverse_transform(self, data, axis=None):
+ _min = paddle.to_tensor(self.min) if paddle.is_tensor(data) else self.min
+ _max = paddle.to_tensor(self.max) if paddle.is_tensor(data) else self.max
+ data = (data + 1.0) / 2.0
+ return 1.0 * data * (_max - _min) + _min
+
+
+class CustomDataset(Dataset):
+ def __init__(self, file_path, data_type="train"):
+ """
+
+ :param file_path:
+ :param data_type: train or test
+ """
+ super().__init__()
+ all_data = np.load(file_path)
+ data = all_data["data"]
+ num, _, _ = data.shape
+ data = data.reshape(num, -1)
+
+ self.neighbors = all_data["neighbors"]
+ self.areasoverlengths = all_data["areasoverlengths"]
+ self.dirichletnodes = all_data["dirichletnodes"]
+ self.dirichleths = all_data["dirichletheads"]
+ self.Qs = np.zeros([all_data["coords"].shape[-1]])
+ self.val_data = all_data["test_data"]
+
+ self.data_type = data_type
+
+ self.train_len = int(num * 0.8)
+ self.test_len = num - self.train_len
+
+ self.train_data = data[: self.train_len]
+ self.test_data = data[self.train_len :]
+
+ self.scaler = ScalerStd()
+ self.scaler.fit(self.train_data)
+
+ self.train_data = self.scaler.transform(self.train_data)
+ self.test_data = self.scaler.transform(self.test_data)
+
+ def __getitem__(self, idx):
+ if self.data_type == "train":
+ return self.train_data[idx]
+ else:
+ return self.test_data[idx]
+
+ def __len__(self):
+ if self.data_type == "train":
+ return self.train_len
+ else:
+ return self.test_len
+
+
+if __name__ == "__main__":
+ train_data = CustomDataset(file_path="data/gaussian_train.npz", data_type="train")
+ test_data = CustomDataset(file_path="data/gaussian_train.npz", data_type="test")
+ train_loader = DataLoader(
+ train_data, batch_size=128, shuffle=True, drop_last=True, num_workers=0
+ )
+ test_loader = DataLoader(
+ test_data, batch_size=128, shuffle=True, drop_last=True, num_workers=0
+ )
+
+ for i, data_item in enumerate(train_loader()):
+ print(data_item)
+
+ if i == 2:
+ break
+
+ np.savez("data.npz", p_train=train_data.train_data, p_test=train_data.test_data)
diff --git a/ppsci/arch/__init__.py b/ppsci/arch/__init__.py
index 7a8e6c263..21416cd44 100644
--- a/ppsci/arch/__init__.py
+++ b/ppsci/arch/__init__.py
@@ -33,6 +33,7 @@
from ppsci.arch.unetex import UNetEx # isort:skip
from ppsci.arch.epnn import Epnn # isort:skip
from ppsci.arch.nowcastnet import NowcastNet # isort:skip
+from ppsci.arch.vae import AutoEncoder # isort:skip
from ppsci.utils import logger # isort:skip
@@ -54,6 +55,7 @@
"UNetEx",
"Epnn",
"NowcastNet",
+ "AutoEncoder",
"build_model",
]
diff --git a/ppsci/arch/vae.py b/ppsci/arch/vae.py
new file mode 100644
index 000000000..202a59d21
--- /dev/null
+++ b/ppsci/arch/vae.py
@@ -0,0 +1,74 @@
+# Copyright (c) 2023 PaddlePaddle Authors. All Rights Reserved.
+
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in compliance with the License.
+# You may obtain a copy of the License at
+
+# http://www.apache.org/licenses/LICENSE-2.0
+
+# Unless required by applicable law or agreed to in writing, software
+# distributed under the License is distributed on an "AS IS" BASIS,
+# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
+# See the License for the specific language governing permissions and
+# limitations under the License.
+
+from __future__ import annotations
+
+from typing import Tuple
+
+import paddle
+import paddle.nn as nn
+
+from ppsci.arch import base
+
+
+# copy from AISTUDIO
+class AutoEncoder(base.Arch):
+ def __init__(
+ self,
+ input_keys: Tuple[str, ...],
+ output_keys: Tuple[str, ...],
+ input_dim,
+ latent_dim,
+ hidden_dim,
+ ):
+ super(AutoEncoder, self).__init__()
+ self.input_keys = input_keys
+ self.output_keys = output_keys
+ # encoder
+ self._encoder_linear = nn.Sequential(
+ nn.Linear(input_dim, hidden_dim),
+ nn.Tanh(),
+ )
+ self._encoder_mu = nn.Linear(hidden_dim, latent_dim)
+ self._encoder_log_sigma = nn.Linear(hidden_dim, latent_dim)
+
+ self._decoder = nn.Sequential(
+ nn.Linear(latent_dim, hidden_dim),
+ nn.Tanh(),
+ nn.Linear(hidden_dim, input_dim),
+ )
+
+ def encoder(self, x):
+ h = self._encoder_linear(x)
+ mu = self._encoder_mu(h)
+ log_sigma = self._encoder_log_sigma(h)
+ return mu, log_sigma
+
+ def decoder(self, x):
+ return self._decoder(x)
+
+ def forward_tensor(self, x):
+ mu, log_sigma = self.encoder(x)
+ z = mu + paddle.randn(mu.shape) * paddle.exp(log_sigma)
+ return mu, log_sigma, self.decoder(z)
+
+ def forward(self, x):
+ x = self.concat_to_tensor(x, self.input_keys, axis=-1)
+ mu, log_sigma, decoder_z = self.forward_tensor(x)
+ result_dict = {
+ self.output_keys[0]: mu,
+ self.output_keys[1]: log_sigma,
+ self.output_keys[2]: decoder_z,
+ }
+ return result_dict
diff --git a/ppsci/loss/__init__.py b/ppsci/loss/__init__.py
index 4e7d6242c..cade922dd 100644
--- a/ppsci/loss/__init__.py
+++ b/ppsci/loss/__init__.py
@@ -17,6 +17,7 @@
from ppsci.loss.base import Loss
from ppsci.loss.func import FunctionalLoss
from ppsci.loss.integral import IntegralLoss
+from ppsci.loss.kl import KLLoss
from ppsci.loss.l1 import L1Loss
from ppsci.loss.l1 import PeriodicL1Loss
from ppsci.loss.l2 import L2Loss
@@ -40,6 +41,7 @@
"MSELoss",
"MSELossWithL2Decay",
"PeriodicMSELoss",
+ "KLLoss",
]
diff --git a/ppsci/loss/kl.py b/ppsci/loss/kl.py
new file mode 100644
index 000000000..6b7143d0d
--- /dev/null
+++ b/ppsci/loss/kl.py
@@ -0,0 +1,45 @@
+# Copyright (c) 2023 PaddlePaddle Authors. All Rights Reserved.
+
+# Licensed under the Apache License, Version 2.0 (the "License");
+# you may not use this file except in compliance with the License.
+# You may obtain a copy of the License at
+
+# http://www.apache.org/licenses/LICENSE-2.0
+
+# Unless required by applicable law or agreed to in writing, software
+# distributed under the License is distributed on an "AS IS" BASIS,
+# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
+# See the License for the specific language governing permissions and
+# limitations under the License.
+
+from __future__ import annotations
+
+from typing import Dict
+from typing import Optional
+from typing import Union
+
+import paddle
+from typing_extensions import Literal
+
+from ppsci.loss import base
+
+
+class KLLoss(base.Loss):
+ def __init__(
+ self,
+ reduction: Literal["mean", "sum"] = "mean",
+ weight: Optional[Union[float, Dict[str, float]]] = None,
+ ):
+ if reduction not in ["mean", "sum"]:
+ raise ValueError(
+ f"reduction should be 'mean' or 'sum', but got {reduction}"
+ )
+ super().__init__(reduction, weight)
+
+ def forward(self, output_dict, label_dict=None, weight_dict=None):
+ mu, log_sigma = output_dict["mu"], output_dict["log_sigma"]
+
+ base = paddle.exp(2.0 * log_sigma) + paddle.pow(mu, 2) - 1.0 - 2.0 * log_sigma
+ loss = 0.5 * paddle.sum(base) / mu.shape[0]
+
+ return loss
From 23f45870a1701bb0b5ad2d6ce84b11413fde7233 Mon Sep 17 00:00:00 2001
From: xusuyong <2209245477@qq.com>
Date: Mon, 22 Jan 2024 11:38:27 +0800
Subject: [PATCH 2/2] add RegAE
---
docs/zh/api/arch.md | 1 +
docs/zh/examples/RegAE.md | 45 ++++++++++++++-------------
examples/RegAE/RegAE.py | 55 +++++++++++++++-----------------
examples/RegAE/conf/RegAE.yaml | 14 ++++-----
examples/RegAE/dataloader.py | 57 +++++++++++++++++++++-------------
ppsci/arch/vae.py | 37 +++++++++++++++++++---
6 files changed, 125 insertions(+), 84 deletions(-)
diff --git a/docs/zh/api/arch.md b/docs/zh/api/arch.md
index 69ff3ac12..32ee405c6 100644
--- a/docs/zh/api/arch.md
+++ b/docs/zh/api/arch.md
@@ -22,5 +22,6 @@
- USCNN
- NowcastNet
- HEDeepONets
+ - AutoEncoder
show_root_heading: true
heading_level: 3
diff --git a/docs/zh/examples/RegAE.md b/docs/zh/examples/RegAE.md
index bbd874c0f..7d41d04eb 100644
--- a/docs/zh/examples/RegAE.md
+++ b/docs/zh/examples/RegAE.md
@@ -1,10 +1,11 @@
-# Learning to regularize with a variational autoencoder for hydrologic inverse analysis
+# Learning to regularize with a variational autoencoder for hydrologic inverse analysis
## 1.简介
本项目基于paddle框架复现。
论文主要点如下:
+
* 作者提出了一种基于变分自动编码器 (VAE)的正则化方法;
* 这种方法的优点1: 对来自VAE的潜在变量(此处指encoder的输出)执行正则化,可以简单地对其进行正则化;
* 这种方法的优点2: VAE减少了优化问题中的变量数量,从而在伴随方法不可用时使基于梯度的优化在计算上更加高效。
@@ -14,8 +15,8 @@
* 实现paddle和julia混写代码梯度传递,避免大面积重写julia代码并能够调用优秀的julia代码;
* 发现paddle minimize_lbfgs存在问题, 待提交issue确认。
-
实验结果要点:
+
* 成功复现论文代码框架及全流程运行测试;
* 本次复现精度因无法使用相同样本,无法与论文中数据进行一一比较。本项目给出了采用paddle编写的框架结果。
@@ -23,16 +24,13 @@
O'Malley D, Golden J K, Vesselinov V V. Learning to regularize with a variational autoencoder for hydrologic inverse analysis[J]. arXiv preprint arXiv:1906.02401, 2019.
参考GitHub地址:
-https://github.com/madsjulia/RegAE.jl
+
项目aistudio地址:
-https://aistudio.baidu.com/aistudio/projectdetail/5541961
+
模型结构
-
-
-
-
+
## 2.数据集
@@ -86,7 +84,6 @@ end
* 数据标准化方式: $z = (x - \mu)/ \sigma$
-
```python
class ScalerStd(object):
"""
@@ -121,6 +118,7 @@ end
3. 数据划分为8:2用与train和test
4. 通过对train数据得到标准化参数mean和std,并用此参数标准化train和test数据集
5. 通过dataloader得到的数据形式为 [batch_size, 10000]
+
```python
class CustomDataset(Dataset):
def __init__(self, file_path, data_type="train"):
@@ -168,6 +166,7 @@ end
else:
return self.test_len
```
+
## 将数据转换为IterableNPZDataset的形式
```python
@@ -184,13 +183,13 @@ np.savez("data.npz", p_train=train_data.train_data, p_test=train_data.test_data)
* Zygote
### python依赖
+
* paddle
* julia (pip安装)
* matplotlib
本项目已经提供安装后压缩文档,可fork本项目后执行以下代码进行解压安装。
-
```python
# 解压预下载文件和预编译文件
!tar zxf /home/aistudio/opt/curl-7.88.1.tar.gz -C /home/aistudio/opt # curl 预下载文件
@@ -200,7 +199,6 @@ np.savez("data.npz", p_train=train_data.train_data, p_test=train_data.test_data)
!tar zxf /home/aistudio/opt/external-libraries.tgz -C /home/aistudio/opt # pip依赖库文件
```
-
```python
####### 以下指令需要时可参考执行,上述压缩包已经完成以下内容 #######
@@ -249,10 +247,12 @@ from julia import Main
## 4.快速开始
本项目运行分为两个步骤:
+
* (1)训练步骤。通过运行train.ipynb文件,可以得到训练后的模型参数,具体代码请参考train.ipynb文件及其中注释说明;
* (2)测试步骤。通过运行test.ipynb文件,应用训练后的模型参数,对latent domain进行优化。
## 5.代码结构与详细说明
+
```text
├── data #预生成数据文件
│ └── data193595
@@ -290,11 +290,11 @@ from julia import Main
### paddle.incubate.optimizer.functional.minimize_lbfgs 问题
以下为paddle官方minimize_lbfgs API:
+
```python
paddle.incubate.optimizer.functional.minimize_lbfgs(objective_func, initial_position, history_size=100, max_iters=50, tolerance_grad=1e-08, tolerance_change=1e-08, initial_inverse_hessian_estimate=None, line_search_fn='strong_wolfe', max_line_search_iters=50, initial_step_length=1.0, dtype='float32', name=None)
```
-
* (1)参数max_line_search_iters无效。虽然设置了此参数,但是内部没有传递对应参数;
* (2)中wolfe条件1错误。line256处应为`phi_2 >= phi_1`,以下为paddle部分源码。
@@ -308,28 +308,30 @@ paddle.incubate.optimizer.functional.minimize_lbfgs(objective_func, initial_posi
## 6.复现结果
### 不同latent维度对比
-
+
+
通过实验结果可以发现:
+
* (1)不同样本之间存在差距,并不是所有样本都能优化得到良好的latent变量;
* (2)随着模型latent维度的上升,模型效果逐渐提升。
### latent_random和latent_mean对比
+
本项目还增加了latent_random和latent_mean对生成结果的对比。此处对latent_random和latent_mean再次说明:
+
* latent_random:通过paddle.randn生成的高斯噪声得到;
* latent_mean:通过对所有testdata进行encoder结果平均得到。
以下为通过latent_random得到的实验结果
-
+
通过对比,可以发现latent_mean对优化结果重要影响。近似正确的latent变量能够得到更优的生成结果。
### LBFGS优化收敛情况
-可以从如下图中看出,使用paddle minimize_lbfgs能够有效优化收敛。
-
-
-
+可以从如下图中看出,使用paddle minimize_lbfgs能够有效优化收敛。
+
## 7.延伸思考
@@ -338,11 +340,10 @@ paddle.incubate.optimizer.functional.minimize_lbfgs(objective_func, initial_posi
回答是肯定的!有意义!
以下为本人个人观点:
+
* (1)通过实验对比latent_random和latent_mean的最终生成结果差距,可以发现一个良好的初值对模型的影响是巨大的。当前diffusion模型在sample生成过程对生成的高斯噪声不断做denoise操作,这其中生成的噪声数据如果经过预先优化,不仅能够加速diffusion的生成速度,而且能够提升数据的生成质量。
* (2)在域迁移等研究领域,可以使用这种latent逐渐生成中间过渡变量,达到不同域数据的迁移生成。
-
-
## 7.模型信息
| 信息 | 说明|
@@ -353,5 +354,5 @@ paddle.incubate.optimizer.functional.minimize_lbfgs(objective_func, initial_posi
| 支持硬件 | GPU、CPU |
| aistudio | [notebook](https://aistudio.baidu.com/aistudio/projectdetail/5541961) |
-请点击[此处](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576)查看本环境基本用法.
-Please click [here ](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576) for more detailed instructions.
+请点击[此处](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576)查看本环境基本用法.
+Please click [here](https://ai.baidu.com/docs#/AIStudio_Project_Notebook/a38e5576) for more detailed instructions.
diff --git a/examples/RegAE/RegAE.py b/examples/RegAE/RegAE.py
index 3170429a9..b32755bde 100644
--- a/examples/RegAE/RegAE.py
+++ b/examples/RegAE/RegAE.py
@@ -17,21 +17,13 @@
from os import path as osp
import hydra
+import paddle
from omegaconf import DictConfig
-from paddle import nn
+from paddle.nn import functional as F
import ppsci
-from ppsci.loss import KLLoss
from ppsci.utils import logger
-criterion = nn.MSELoss()
-kl_loss = KLLoss()
-
-
-def loss_expr(output_dict, label_dict, weight_dict=None):
-
- return kl_loss(output_dict) + criterion(output_dict["p_hat"], label_dict["p_hat"])
-
def train(cfg: DictConfig):
# set random seed for reproducibility
@@ -45,24 +37,30 @@ def train(cfg: DictConfig):
# set dataloader config
train_dataloader_cfg = {
"dataset": {
- "name": "IterableNPZDataset",
+ "name": "NPZDataset",
"file_path": cfg.TRAIN_FILE_PATH,
- "input_keys": ("p",),
- "label_keys": ("p_hat",),
- "alias_dict": {"p": "p_train", "p_hat": "p_train"},
+ "input_keys": ("p_train",),
+ "label_keys": ("p_train",),
},
"batch_size": cfg.TRAIN.batch_size,
"sampler": {
"name": "BatchSampler",
- "drop_last": False,
- "shuffle": True,
+ "drop_last": True,
+ "shuffle": False,
},
}
+ def loss_expr(output_dict, label_dict, weight_dict=None):
+ mu, log_sigma = output_dict["mu"], output_dict["log_sigma"]
+
+ base = paddle.exp(2.0 * log_sigma) + paddle.pow(mu, 2) - 1.0 - 2.0 * log_sigma
+ KLLoss = 0.5 * paddle.sum(base) / mu.shape[0]
+
+ return F.mse_loss(output_dict["decoder_z"], label_dict["p_train"]) + KLLoss
+
# set constraint
sup_constraint = ppsci.constraint.SupervisedConstraint(
train_dataloader_cfg,
- output_expr={"p_hat": lambda out: out["p_hat"]},
loss=ppsci.loss.FunctionalLoss(loss_expr),
name="Sup",
)
@@ -75,23 +73,21 @@ def train(cfg: DictConfig):
# set validator
eval_dataloader_cfg = {
"dataset": {
- "name": "IterableNPZDataset",
+ "name": "NPZDataset",
"file_path": cfg.VALID_FILE_PATH,
- "input_keys": ("p",),
- "label_keys": ("p_hat",),
- "alias_dict": {"p": "p_train", "p_hat": "p_train"},
+ "input_keys": ("p_train",),
+ "label_keys": ("p_train",),
},
"batch_size": cfg.EVAL.batch_size,
"sampler": {
"name": "BatchSampler",
- "drop_last": False,
+ "drop_last": True,
"shuffle": False,
},
}
sup_validator = ppsci.validate.SupervisedValidator(
eval_dataloader_cfg,
- loss=ppsci.loss.MSELoss(),
- output_expr={"p_hat": lambda out: out["p_hat"]},
+ loss=ppsci.loss.FunctionalLoss(loss_expr),
metric={"L2Rel": ppsci.metric.L2Rel()},
)
validator = {sup_validator.name: sup_validator}
@@ -121,7 +117,7 @@ def evaluate(cfg: DictConfig):
# set random seed for reproducibility
ppsci.utils.misc.set_random_seed(cfg.seed)
# initialize logger
- logger.init_logger("ppsci", osp.join(cfg.output_dir, "train.log"), "info")
+ logger.init_logger("ppsci", osp.join(cfg.output_dir, "eval.log"), "info")
# set model
model = ppsci.arch.AutoEncoder(**cfg.MODEL)
@@ -129,16 +125,15 @@ def evaluate(cfg: DictConfig):
# set validator
eval_dataloader_cfg = {
"dataset": {
- "name": "IterableNPZDataset",
+ "name": "NPZDataset",
"file_path": cfg.VALID_FILE_PATH,
- "input_keys": ("p",),
- "label_keys": ("p_hat",),
- "alias_dict": {"p": "p_train", "p_hat": "p_train"},
+ "input_keys": ("p_train",),
+ "label_keys": ("p_train",),
},
"batch_size": cfg.EVAL.batch_size,
"sampler": {
"name": "BatchSampler",
- "drop_last": False,
+ "drop_last": True,
"shuffle": False,
},
}
diff --git a/examples/RegAE/conf/RegAE.yaml b/examples/RegAE/conf/RegAE.yaml
index 133ac82a5..15c57f82a 100644
--- a/examples/RegAE/conf/RegAE.yaml
+++ b/examples/RegAE/conf/RegAE.yaml
@@ -21,24 +21,24 @@ hydra:
# general settings
mode: train # running mode: train/eval
-seed: 42
+seed: 1
output_dir: ${hydra:run.dir}
TRAIN_FILE_PATH: data.npz
VALID_FILE_PATH: data.npz
# model settings
MODEL:
- input_keys: [ "p",]
- output_keys: ["mu", "log_sigma", "p_hat"]
+ input_keys: ["p_train",]
+ output_keys: ["mu", "log_sigma", "decoder_z"]
input_dim: 10000
latent_dim: 100
hidden_dim: 100
# training settings
TRAIN:
- epochs: 200
- iters_per_epoch: 1
- eval_during_train: true
+ epochs: 2
+ iters_per_epoch: 625
+ eval_during_train: False
save_freq: 200
eval_freq: 200
learning_rate: 0.0001
@@ -47,5 +47,5 @@ TRAIN:
# evaluation settings
EVAL:
pretrained_model_path: null
- eval_with_no_grad: False
+ eval_with_no_grad: True
batch_size: 128
diff --git a/examples/RegAE/dataloader.py b/examples/RegAE/dataloader.py
index e64654b96..dbeb23611 100644
--- a/examples/RegAE/dataloader.py
+++ b/examples/RegAE/dataloader.py
@@ -1,16 +1,15 @@
"""
-输入数据类型 10^5 * 100 * 100
+输入数据形状 10^5 * 100 * 100
1.按照8:2划分训练数据集和测试数据集
2.通过训练数据进行标准正则化
"""
import numpy as np
import paddle
-from paddle.io import DataLoader
-from paddle.io import Dataset
+from paddle import io
-class ScalerStd(object):
+class ZScoreNormalize:
"""
Desc: Normalization utilities with std mean
"""
@@ -25,24 +24,32 @@ def fit(self, data):
def transform(self, data):
mean = (
- paddle.to_tensor(self.mean).type_as(data).to(data.device)
+ paddle.to_tensor(self.mean, dtype=data.dtype)
if paddle.is_tensor(data)
else self.mean
)
std = (
- paddle.to_tensor(self.std).type_as(data).to(data.device)
+ paddle.to_tensor(self.std, dtype=data.dtype)
if paddle.is_tensor(data)
else self.std
)
return (data - mean) / std
def inverse_transform(self, data):
- mean = paddle.to_tensor(self.mean) if paddle.is_tensor(data) else self.mean
- std = paddle.to_tensor(self.std) if paddle.is_tensor(data) else self.std
+ mean = (
+ paddle.to_tensor(self.mean, dtype=data.dtype)
+ if paddle.is_tensor(data)
+ else self.mean
+ )
+ std = (
+ paddle.to_tensor(self.std, dtype=data.dtype)
+ if paddle.is_tensor(data)
+ else self.std
+ )
return (data * std) + mean
-class ScalerMinMax(object):
+class MinMaxNormalize:
"""
Desc: Normalization utilities with min max
"""
@@ -57,12 +64,12 @@ def fit(self, data):
def transform(self, data):
_min = (
- paddle.to_tensor(self.min).type_as(data).to(data.device)
+ paddle.to_tensor(self.min, dtype=data.dtype)
if paddle.is_tensor(data)
else self.min
)
_max = (
- paddle.to_tensor(self.max).type_as(data).to(data.device)
+ paddle.to_tensor(self.max, dtype=data.dtype)
if paddle.is_tensor(data)
else self.max
)
@@ -70,13 +77,21 @@ def transform(self, data):
return 2.0 * data - 1.0
def inverse_transform(self, data, axis=None):
- _min = paddle.to_tensor(self.min) if paddle.is_tensor(data) else self.min
- _max = paddle.to_tensor(self.max) if paddle.is_tensor(data) else self.max
+ _min = (
+ paddle.to_tensor(self.min, dtype=data.dtype)
+ if paddle.is_tensor(data)
+ else self.min
+ )
+ _max = (
+ paddle.to_tensor(self.max, dtype=data.dtype)
+ if paddle.is_tensor(data)
+ else self.max
+ )
data = (data + 1.0) / 2.0
return 1.0 * data * (_max - _min) + _min
-class CustomDataset(Dataset):
+class CustomDataset(io.Dataset):
def __init__(self, file_path, data_type="train"):
"""
@@ -104,11 +119,11 @@ def __init__(self, file_path, data_type="train"):
self.train_data = data[: self.train_len]
self.test_data = data[self.train_len :]
- self.scaler = ScalerStd()
- self.scaler.fit(self.train_data)
+ self.normalizer = ZScoreNormalize()
+ self.normalizer.fit(self.train_data)
- self.train_data = self.scaler.transform(self.train_data)
- self.test_data = self.scaler.transform(self.test_data)
+ self.train_data = self.normalizer.transform(self.train_data)
+ self.test_data = self.normalizer.transform(self.test_data)
def __getitem__(self, idx):
if self.data_type == "train":
@@ -126,10 +141,10 @@ def __len__(self):
if __name__ == "__main__":
train_data = CustomDataset(file_path="data/gaussian_train.npz", data_type="train")
test_data = CustomDataset(file_path="data/gaussian_train.npz", data_type="test")
- train_loader = DataLoader(
+ train_loader = io.DataLoader(
train_data, batch_size=128, shuffle=True, drop_last=True, num_workers=0
)
- test_loader = DataLoader(
+ test_loader = io.DataLoader(
test_data, batch_size=128, shuffle=True, drop_last=True, num_workers=0
)
@@ -139,4 +154,4 @@ def __len__(self):
if i == 2:
break
- np.savez("data.npz", p_train=train_data.train_data, p_test=train_data.test_data)
+ # np.savez("data.npz", p_train=train_data.train_data, p_test=train_data.test_data)
diff --git a/ppsci/arch/vae.py b/ppsci/arch/vae.py
index 202a59d21..2a05f0d64 100644
--- a/ppsci/arch/vae.py
+++ b/ppsci/arch/vae.py
@@ -22,15 +22,44 @@
from ppsci.arch import base
-# copy from AISTUDIO
class AutoEncoder(base.Arch):
+ """
+ AutoEncoder is a class that represents an autoencoder neural network model.
+
+ Args:
+ input_keys (Tuple[str, ...]): A tuple of input keys.
+ output_keys (Tuple[str, ...]): A tuple of output keys.
+ input_dim (int): The dimension of the input data.
+ latent_dim (int): The dimension of the latent space.
+ hidden_dim (int): The dimension of the hidden layer.
+
+ Examples:
+ >>> import paddle
+ >>> import ppsci
+ >>> model = ppsci.arch.AutoEncoder(
+ ... input_keys=("input1",),
+ ... output_keys=("mu", "log_sigma", "decoder_z",),
+ ... input_dim=100,
+ ... latent_dim=50,
+ ... hidden_dim=200
+ ... )
+ >>> input_dict = {"input1": paddle.rand([200, 100]),}
+ >>> output_dict = model(input_dict)
+ >>> print(output_dict["mu"].shape)
+ [200, 50]
+ >>> print(output_dict["log_sigma"].shape)
+ [200, 50]
+ >>> print(output_dict["decoder_z"].shape)
+ [200, 100]
+ """
+
def __init__(
self,
input_keys: Tuple[str, ...],
output_keys: Tuple[str, ...],
- input_dim,
- latent_dim,
- hidden_dim,
+ input_dim: int,
+ latent_dim: int,
+ hidden_dim: int,
):
super(AutoEncoder, self).__init__()
self.input_keys = input_keys